Gerber:What We Do: Difference between revisions
From OpenWetWare
Jump to navigationJump to search
No edit summary |
No edit summary |
||
| Line 1: | Line 1: | ||
{{Template:Gerber}} | {{Template:Gerber}} | ||
====Global analysis of post-transcriptional gene regulation==== | |||
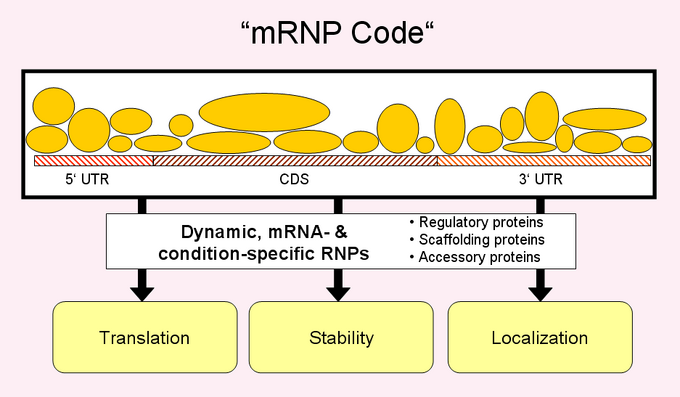
Post-transcriptional control of gene expression involves RNA processing, transport, turnover and mRNA translation. Regulation of these steps has substantial effects on the expression and function of genes in diverse processes such cytokinesis, embryonic development, neurogenesis and cancer progression. RNA-binding proteins (RBPs) have been implicated in diverse aspects of post-transcriptional gene expression. Hundreds of RBPs are encoded in eukaryotic genomes, and whereas many classical studies explored the cellular role of RBPs with specific mRNA substrates, the recent development of genome-wide analysis tools enables systematic identification of mRNA substrates of RBPs, and the study of post-transcriptional gene regulation on a global scale. Importantly, these studies revealed that many RBPs bind to and regulate subsets of mRNAs, which encode proteins that are localized to the same subcellular compartment, act in the same pathway or are components of macromolecular complexes. Moreover, these sets of mRNAs often contain characteristic sequence elements in the 3'-untranslated regions, which are potential binding sites for regulatory RBPs (Gerber et al. 2004). These findings strongly indicate the presence of extensive post-transcriptional regulatory systems in eukaryotic cells, which may be comparable in its extent and richness to that of transcriptional regulatory networks. | Post-transcriptional control of gene expression involves RNA processing, transport, turnover and mRNA translation. Regulation of these steps has substantial effects on the expression and function of genes in diverse processes such cytokinesis, embryonic development, neurogenesis and cancer progression. RNA-binding proteins (RBPs) have been implicated in diverse aspects of post-transcriptional gene expression. Hundreds of RBPs are encoded in eukaryotic genomes, and whereas many classical studies explored the cellular role of RBPs with specific mRNA substrates, the recent development of genome-wide analysis tools enables systematic identification of mRNA substrates of RBPs, and the study of post-transcriptional gene regulation on a global scale. Importantly, these studies revealed that many RBPs bind to and regulate subsets of mRNAs, which encode proteins that are localized to the same subcellular compartment, act in the same pathway or are components of macromolecular complexes. Moreover, these sets of mRNAs often contain characteristic sequence elements in the 3'-untranslated regions, which are potential binding sites for regulatory RBPs (Gerber et al. 2004). These findings strongly indicate the presence of extensive post-transcriptional regulatory systems in eukaryotic cells, which may be comparable in its extent and richness to that of transcriptional regulatory networks. | ||