IGEM:MIT/2006/Notebook/2006-7-19: Difference between revisions
| (3 intermediate revisions by one other user not shown) | |||
| Line 65: | Line 65: | ||
.5 uL EcoRI | .5 uL EcoRI | ||
===Ran 2 gels=== | |||
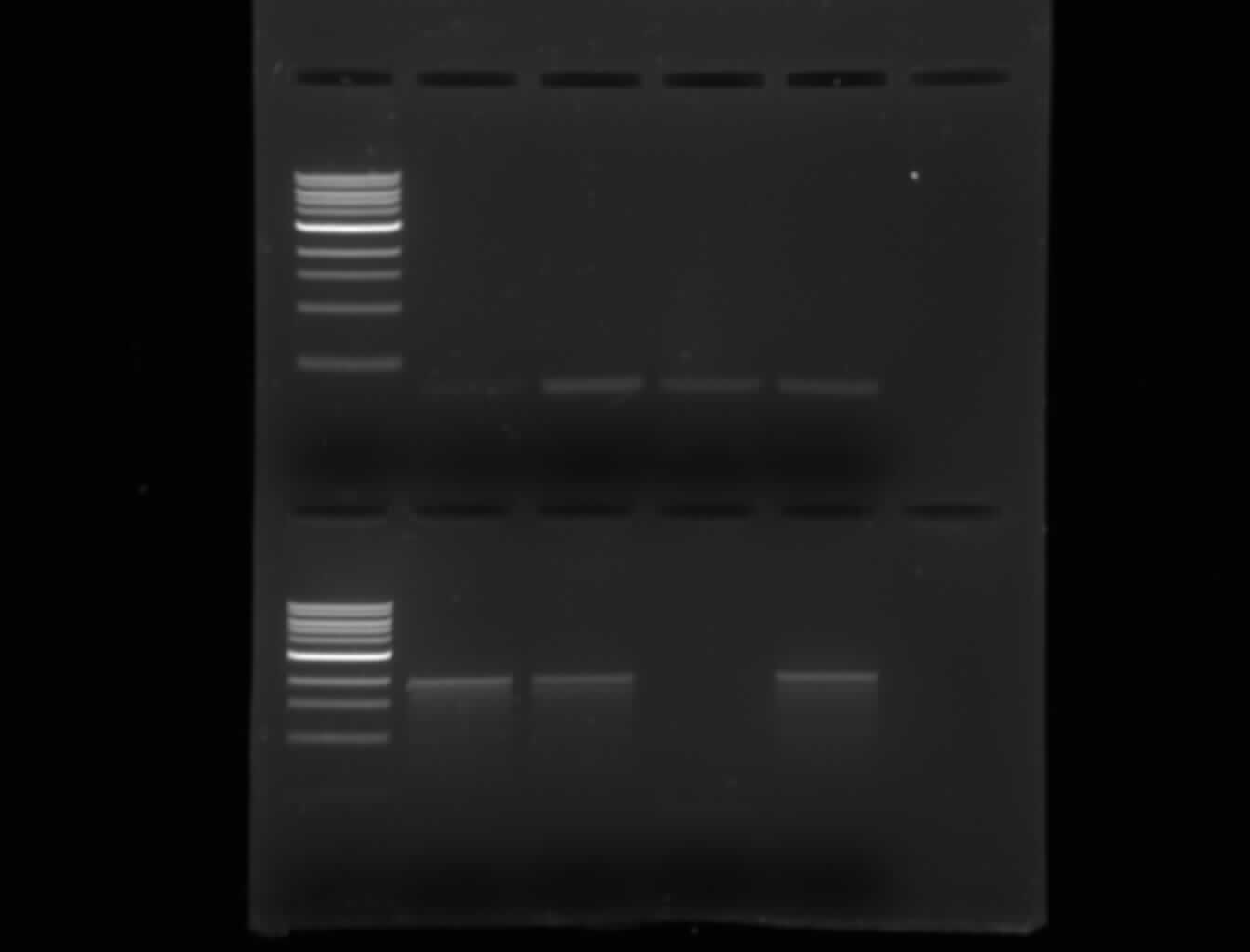
...nothing showed up on the first gel, which was the same as the gel below, but it had RBS30 instead of 32, ATF-A instead of ATF-B, and no promoter: | |||
Lanes: | |||
*1kB ladder | |||
*2-log ladder | |||
*RBS 32 / SAMT / term 15 in pSB1A3-1 cut with EcoRI and XbaI [from the SAMT+terminator colony A] | |||
*RBS 32 / SAMT / term 15 in pSB1A3-1 cut with EcoRI and XbaI [from the SAMT+terminator colony B] | |||
*RBS 32 / SAMT / term 15 in pSB1A3-1 cut with EcoRI and XbaI [from the SAMT+terminator colony C] | |||
*RBS 32 / SAMT / term 15 in pSB1A3-1 cut with EcoRI and XbaI [from the SAMT+terminator colony D] | |||
*ATF1-B, mutagenized, cut with EcoRI to see if the mutagenesis worked. | |||
*Promoter: R0040, cut with EcoRI and SpeI | |||
[[Image:7-19.jpg]] | |||
==Ligation== | ==Ligation== | ||
| Line 70: | Line 84: | ||
What we need to do is ligate R0040 to the SAMT.B0015 constructs that we have. | What we need to do is ligate R0040 to the SAMT.B0015 constructs that we have. | ||
1.5μL H20 | |||
3.5μLL RBS (15-25 ng/uL) | 3.5μLL RBS (15-25 ng/uL) | ||
3.5μLL Enzyme+B.0015 (15-25 ng/uL) | 3.5μLL Enzyme+B.0015 (15-25 ng/uL) | ||
1μLL T4 DNA Ligase Buffer | 1μLL T4 DNA Ligase Buffer | ||
0.5μLL T4 DNA Ligase | 0.5μLL T4 DNA Ligase | ||
==Transformation== | |||
The following ligation mixes were transformed: | |||
R0040.B0030.SAMT.B0015 A, B, C, & D | |||
R0040.B0032.SAMT.B0015 A, B, C, & D | |||
Latest revision as of 14:08, 20 July 2006
Smell Cultures
- 4 BSMT (2 SA, 2 BA) and 2 BAMT (ctrl not induced)
Sequencing Results
Positive for SAMT.B0015 C! The one we used!
Miniprep & Subsequent Steps Taken
We miniprepped each liquid culture from last night. We found out that SAMTA.B0015 was incorrect and decided to discard B0030.SAMTA.B0015 and B0032.SAMTA.B0015. We also diluted 20 uL of the indole knockout strains containing BAMT and BSMT into 10 mL of liquid culture and 40 uL of 1 M Benzoic Acid (it actually probably should have been 20 uL). We then sequenced each of the remaining minipreps and digested each one. Note that we decided to digest ATF1 mutagenesis colonies A and B with EcoRI, the site that should have been mutated in site-directed mutagenesis.
Digests
R0040
27.5 uL H20
5 uL Buffer 2
.5 uL BSA
16 uL 50 ng/uL DNA
.5 uL EcoRI
.5 uL SpeI
B0030.SAMT.B0015
27.5 uL H20
5 uL Buffer 2
.5 uL BSA
16 uL 50 ng/uL DNA
.5 uL EcoRI
.5 uL XbaI
B0030.SAMT.B0015 A, B, & C + B0032.SAMT.B0015 A, B, C, & D
35.5 uL H20
5 uL Buffer 2
.5 uL BSA
8 uL 100 ng/uL DNA
.5 uL EcoRI
.5 uL XbaI
ATF1 Mutagenesis A & B
36 uL H20
5 uL Buffer 2
.5 uL BSA
8 uL 100 ng/uL DNA
.5 uL EcoRI
Ran 2 gels
...nothing showed up on the first gel, which was the same as the gel below, but it had RBS30 instead of 32, ATF-A instead of ATF-B, and no promoter:
Lanes:
- 1kB ladder
- 2-log ladder
- RBS 32 / SAMT / term 15 in pSB1A3-1 cut with EcoRI and XbaI [from the SAMT+terminator colony A]
- RBS 32 / SAMT / term 15 in pSB1A3-1 cut with EcoRI and XbaI [from the SAMT+terminator colony B]
- RBS 32 / SAMT / term 15 in pSB1A3-1 cut with EcoRI and XbaI [from the SAMT+terminator colony C]
- RBS 32 / SAMT / term 15 in pSB1A3-1 cut with EcoRI and XbaI [from the SAMT+terminator colony D]
- ATF1-B, mutagenized, cut with EcoRI to see if the mutagenesis worked.
- Promoter: R0040, cut with EcoRI and SpeI
Ligation
What we need to do is ligate R0040 to the SAMT.B0015 constructs that we have.
1.5μL H20
3.5μLL RBS (15-25 ng/uL)
3.5μLL Enzyme+B.0015 (15-25 ng/uL)
1μLL T4 DNA Ligase Buffer
0.5μLL T4 DNA Ligase
Transformation
The following ligation mixes were transformed:
R0040.B0030.SAMT.B0015 A, B, C, & D
R0040.B0032.SAMT.B0015 A, B, C, & D