BME494 Sp2014 Dhatt: Difference between revisions
From OpenWetWare
Jump to navigationJump to search
No edit summary |
No edit summary |
||
| Line 49: | Line 49: | ||
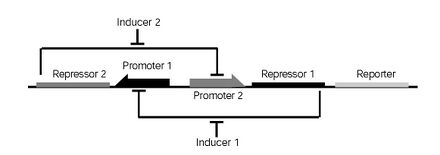
<!-- Show a network/ circuit diagram of your device. Include a paragraph to explain how it works (e.g., how to switch the system from on to off and vice versa, and what happens to each component as the system switches between states) --> | <!-- Show a network/ circuit diagram of your device. Include a paragraph to explain how it works (e.g., how to switch the system from on to off and vice versa, and what happens to each component as the system switches between states) --> | ||
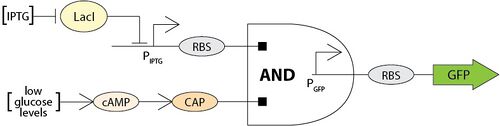
[[Image:BME494_Dhatt_AndGate.jpg| | [[Image:BME494_Dhatt_AndGate.jpg|500px|center|AND gate logic gene toggle switch. IPTG and low glucose levels conditions must be met in order for GFP production]] | ||
<br>AND gate logic gene toggle switch. IPTG and low glucose levels conditions must be met in order for GFP production.<br> | <br>AND gate logic gene toggle switch. IPTG and low glucose levels conditions must be met in order for GFP production.<br> | ||
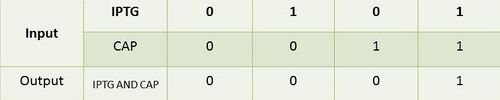
[[Image:BME494_Dhatt_AndGateTable.jpg| | [[Image:BME494_Dhatt_AndGateTable.jpg|500px|center|Table describing that both inputs are needed in order to produce an output]] | ||
<br>Table describing that both inputs are needed in order to produce an output.<br> | <br>Table describing that both inputs are needed in order to produce an output.<br> | ||
| Line 64: | Line 64: | ||
'''SYNTHETIC DNA LAYOUT''' | '''SYNTHETIC DNA LAYOUT''' | ||
Type IIs Assembly was used to build this lac switch. Type IIs Assembly allows for parts to be assembled in one step. For Type IIs Assembly, forward and reverse primers are needed to be created and placed in the system in order to create sticky ends that can bind various parts together. To put the pieces together, PCR is implemented, which allows all the parts to be replicated thousands of times in order to produce a desired final product. Digestion and ligation is used during which BsmBI cuts the DNA fragments and creates complementary overhangs that anneal via base pairing. | |||
<!-- Show each DNA fragment in the order in which you propose to assemble them into a single plasmid. This can be a series of connected blocks. You do not have to show the plasmid as a circle. --> | <!-- Show each DNA fragment in the order in which you propose to assemble them into a single plasmid. This can be a series of connected blocks. You do not have to show the plasmid as a circle. --> | ||
'''RESOURCES''' | '''RESOURCES''' | ||
<!-- List pre-existing parts/ Biobricks that you will use. Also list new parts that you will need to get from a natural source. BE SPECIFIC. --> | <!-- List pre-existing parts/ Biobricks that you will use. Also list new parts that you will need to get from a natural source. BE SPECIFIC. --> | ||
The following BioBrick parts, found on the iGEM registry website, will be used to build the new device: | |||
K418003 - composite Lac promoter inducible by IPTG | |||
• Size: 1416 base pairs | |||
• LacI present: transcription inhibited | |||
• LacI absent: transcription promoted | |||
• LacI inhibited by IPTG | |||
B0030 – Strong RBS | |||
• Size: 15 base pairs | |||
K259006 – Composite part made from green fluorescent protein (GFP) and double terminator | |||
• IPTG present: GFP produced | |||
• IPTG absent: GFP not produced, clear solution | |||
• Size: 823 base pairs | |||
pSB1A3 – BioBrick assembly plasmid | |||
• Size:2155 base pairs | |||
'''TYPE IIS ASSEMBLY''' | '''TYPE IIS ASSEMBLY''' | ||
'''PCR''' | |||
Polymerase Chain Reaction (PCR) will be used for amplification of the DNA parts. PCR is the process of adding DNA, primers, nucleotides and DNA polymerase to a tube which produces system through assembly pot process after placed in a thermocycler and placed through many cycles. PCR consists of three specific temperature steps: denaturation, annealing, and elongation. Denaturation allows the DNA template to be heated to a specific temperature and yield single-stranded DNA molecules. The reaction temperature is then lowered for a short period of time in order to allow the complementary DNA primers to anneal to the single-stranded DNA template. The primers are then elongated by the DNA polymerase that is used. The DNA polymerase then allows for the cycle to be repeated multiple times resulting in thousands of copies of the desired fragments. | |||
<br> | |||
'''Digestion/Ligation Reaction''' | |||
The digestion/ligation reaction allows for dilution the purified PCR products to volume of 20 ul. The digestion/ligation reaction uses BsmB1, a Type II restriction enzyme, which cuts DNA fragments to create complementary overhangs (sticky ends). These sticky ends anneal via DNA base pairs. Primers introduce these complementary overhangs to the DNA. T4 ligase is used to seal the gaps between base pairs and create a finished system. | |||
<br> | |||
'''Primers''' | |||
The following primers will be used for the new device in order to introduce complementary overhangs and BsmBI cut sites. | |||
Forward Primer Vector: 5'-cacaccaCGTCTCaactagtagcggccgct | |||
Reverse Primer Vector: 5'-cacaccaCGTCTCatctagatgcggccgcg | |||
Forward Primer Composite Lac Promoter: 5'-cacaccaCGTCTCatagattgacagctagctca | |||
Reverse Primer Composite Lac Promoter: 5'-cacaccaCGTCTCatgtgtgtgctcagtatctt | |||
Forward Primer Operator: 5'-cacaccaCGTCTCacaatacgcaaaccgc | |||
Reverse Primer Operator: 5'-cacaccaCGTCTCatttctgtgtgaaattgtta | |||
Forward Primer RBS: 5'-cacaccaCGTCTCattaaagaggagaaa | |||
Reverse Primer RBS: 5'-cacaccaCGTCTCaaattttctcctctttaat | |||
Forward Primer GFP with terminator: 5'-cacaccaCGTCTCaatgcgtaaaggagaa | |||
Reverse Primer GFP with terminator: 5'-cacaccaCGTCTCatagtaaataataaaaaagc | |||
Forward Primer for the mutation: 5’-agctgttgccGgtctcactgg | |||
Reverse Primer for the mutation: 5’-ccagtgagacCggcaacagct | |||
<br> | |||
'''Reagents''' | |||
PCR Reagents: | |||
{| border="1" cellpadding="5" style="width: 100%" class="wikitable" | |||
|+ '''PCR Reagents''' | |||
! Reagent | |||
! Volume | |||
|- | |||
! Template DNA | |||
| 0.2 μL | |||
|- | |||
! 1.0 uM Forward Primer | |||
| 1.0 μL | |||
|- | |||
! 1.0 uM Reverse Primer | |||
| 1.0 μL | |||
|- | |||
! 2x GoTaq Green Mix | |||
| 25 μL | |||
|- | |||
! H<sub>2</sub>O | |||
| 22.8 μL | |||
|- | |||
! TOTAL VOLUME | |||
| 50 μL | |||
|- | |||
|}<br> | |||
The PCR reagents used the following thermal cycling properties: <br> | |||
95°C, 3 min.<br> | |||
[95°C, 15 sec; 55°C, 15 sec; 72°C, 30 sec] x30<br> | |||
72°C, 3 min.<br> | |||
4°C, ∞<br> | |||
<!-- Incorporate information from Presentation 2 --> | <!-- Incorporate information from Presentation 2 --> | ||
<br> Digestipn/Ligation Reaction Reagents: | |||
{| border="1" cellpadding="5" style="width: 100%" class="wikitable" | |||
|+ '''Digestion/Ligation Reagents''' | |||
! Reagent | |||
! Assembly | |||
! Negative Control | |||
|- | |||
| PCR fragment 1 (vector) | |||
| 1.0 μL | |||
| 1.0 μL | |||
|- | |||
| PCR fragment 2 | |||
| 1.0 μL | |||
| None | |||
|- | |||
| PCR fragment 3 | |||
| 1.0 μL | |||
| None | |||
|- | |||
| PCR fragment 4 | |||
| 1.0 μL | |||
| None | |||
|- | |||
| PCR fragment 5 | |||
| 1.0 μL | |||
| None | |||
|- | |||
| 10x T4 Ligase | |||
| 0.25 μL | |||
| 0.25 μL | |||
|- | |||
| BsmB1 | |||
| 0.5 μL | |||
| 0.5 μL | |||
|- | |||
| H<sub>2</sub>O | |||
| 4.25 μL | |||
| 8.25 μL | |||
|- | |||
| Total Volume | |||
| 10 μL | |||
| 10 μL | |||
|- | |||
|}<br> | |||
The Digestion/Ligation reagents used the following thermal cycling properties: <br> | |||
[45°C, 2 min.; 16°C, 5 min] x25<br> | |||
60°C, 10 min. | |||
80°C, 20 min.<br> | |||
4°C, ∞<br> | |||
<!-- Incorporate information from Presentation 2 --> | |||
Revision as of 22:30, 8 May 2014
My Profile Dr. Haynes OpenWetWare Previous Course Wiki Editing Help