Maloof Lab:Research: Difference between revisions
From OpenWetWare
Jump to navigationJump to search
mNo edit summary |
mNo edit summary |
||
| Line 10: | Line 10: | ||
Interestingly, the correct response to a specific light cue depends on the environmental context. Thus, plants native to different light environments have evolved different adaptive responses. One clear example is that of shade avoidance. Light that is transmitted through or reflected from neighboring plants is reduced in its red to far-red (R/FR) ratio, because chlorophyll absorbs red but not far-red light. The phytochrome photoreceptors detect this change in light quality and trigger a suite of responses that increase light competiveness, including stem and petiole elongation and early reproduction<sup>3</sup> (Figure 1). | Interestingly, the correct response to a specific light cue depends on the environmental context. Thus, plants native to different light environments have evolved different adaptive responses. One clear example is that of shade avoidance. Light that is transmitted through or reflected from neighboring plants is reduced in its red to far-red (R/FR) ratio, because chlorophyll absorbs red but not far-red light. The phytochrome photoreceptors detect this change in light quality and trigger a suite of responses that increase light competiveness, including stem and petiole elongation and early reproduction<sup>3</sup> (Figure 1). | ||
[[Image:Research_fig_1.jpg | center]] | [[Image:Research_fig_1.jpg | center | Figure 1]] | ||
Plants native to sunny environments are very sensitive to shade from their neighbors and show robust shade avoidance responses, allowing them to compete for sunlight. In contrast, shade avoidance is much reduced in plants native to constitutively shady environments such as forest understory. Such changes in shade avoidance response are adaptive and are due to heritable genetic differences<sup>4-6</sup>. Although mutational studies have defined a number of genes involved in phytochrome signaling<sup>7,8</sup>, the genes and molecular changes responsible for adaptive changes in light response remain unknown. | Plants native to sunny environments are very sensitive to shade from their neighbors and show robust shade avoidance responses, allowing them to compete for sunlight. In contrast, shade avoidance is much reduced in plants native to constitutively shady environments such as forest understory. Such changes in shade avoidance response are adaptive and are due to heritable genetic differences<sup>4-6</sup>. Although mutational studies have defined a number of genes involved in phytochrome signaling<sup>7,8</sup>, the genes and molecular changes responsible for adaptive changes in light response remain unknown. | ||
| Line 17: | Line 17: | ||
==Prior Work== | ==Prior Work== | ||
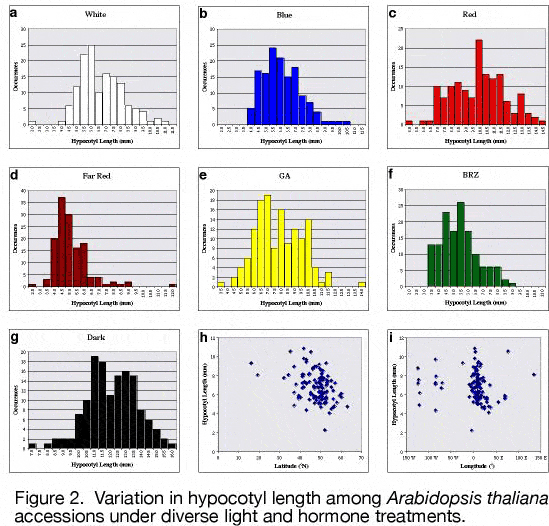
To determine if ''A. thaliana'' is an appropriate organism for study of light adaptation, we characterized the extent of light response variation in 140 ''A. thaliana'' accessions using a simple response, seedling emergence<sup>9</sup>. A wide range of heritable differences in light response was found (Figure 2). | To determine if ''A. thaliana'' is an appropriate organism for study of light adaptation, we characterized the extent of light response variation in 140 ''A. thaliana'' accessions using a simple response, seedling emergence<sup>9</sup>. A wide range of heritable differences in light response was found (Figure 2). | ||
[[Image:Research_fig_2.jpg | center]] | [[Image:Research_fig_2.jpg | center | Figure 2]] | ||
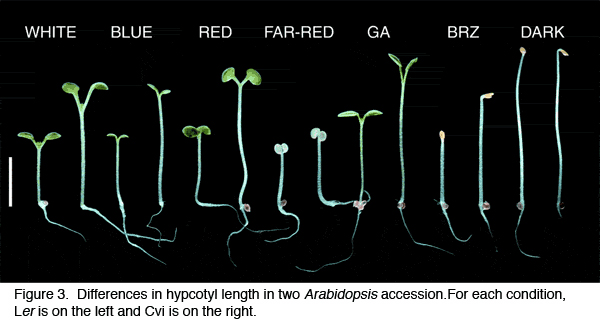
Interestingly, there was a significant inverse correlation between latitude and light sensitivity, suggesting adaptation to an environmental factor that varies over latitudinal clines (perhaps light itself). Some accessions had novel response patterns whereas others had patterns suggesting changes in known pathways. One accession, ''Lm-2'', was insensitive to far-red light, similar to phytochromeA (phyA) mutants. We found that ''Lm-2'' fails to complement phyA due to a single amino acid change and that this change causes production of a protein that is less sensitive to light. Biochemical characterization of ''Lm-2'' phytochrome demonstrated that it has reduced kinase activity and altered spectral properties. | Interestingly, there was a significant inverse correlation between latitude and light sensitivity, suggesting adaptation to an environmental factor that varies over latitudinal clines (perhaps light itself). Some accessions had novel response patterns whereas others had patterns suggesting changes in known pathways. One accession, ''Lm-2'', was insensitive to far-red light, similar to phytochromeA (phyA) mutants. We found that ''Lm-2'' fails to complement phyA due to a single amino acid change and that this change causes production of a protein that is less sensitive to light. Biochemical characterization of ''Lm-2'' phytochrome demonstrated that it has reduced kinase activity and altered spectral properties. | ||
| Line 27: | Line 26: | ||
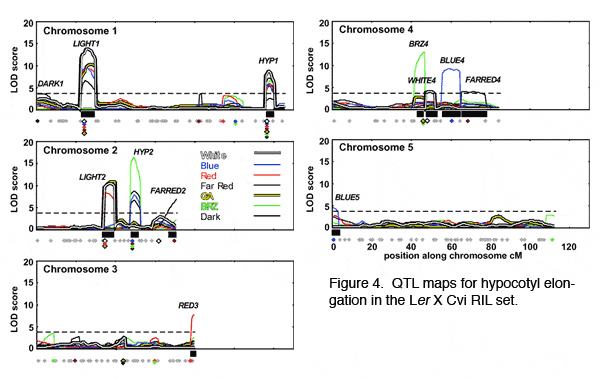
[[Image:Research_fig_3.jpg | center]] | [[Image:Research_fig_3.jpg | center | Figure 3]] | ||
[[Image:Research_fig_4.jpg | center]] | [[Image:Research_fig_4.jpg | center | Figure 4]] | ||
Some loci map to regions with no known photomorphogenic mutants, suggesting that new genes involved in light response have been identified. In contrast, one QTL maps near PHYTOCHROMEB (PHYB), known to be important for response to red light. We created a near-isogenic line (NIL) that confirmed that this region is important for light response, and preliminary results from transgenic plants suggest that PHYB may indeed be the QTL. Strikingly, association testing suggests that the PHYB region is an important determinant of light response across many ''A. thaliana'' accessions. Furthermore, sequence comparison between ''A. thaliana'' and ''Arabidopsis lyrata'' show that PHYB is evolving in a non-neutral fashion. Combined, these studies demonstrate that Arabidopsis is an excellent organism for studying the molecular basis of natural variation in light response, and suggest that some changes in light response in Arabidopsis and its relatives is adaptive. | Some loci map to regions with no known photomorphogenic mutants, suggesting that new genes involved in light response have been identified. In contrast, one QTL maps near PHYTOCHROMEB (PHYB), known to be important for response to red light. We created a near-isogenic line (NIL) that confirmed that this region is important for light response, and preliminary results from transgenic plants suggest that PHYB may indeed be the QTL. Strikingly, association testing suggests that the PHYB region is an important determinant of light response across many ''A. thaliana'' accessions. Furthermore, sequence comparison between ''A. thaliana'' and ''Arabidopsis lyrata'' show that PHYB is evolving in a non-neutral fashion. Combined, these studies demonstrate that Arabidopsis is an excellent organism for studying the molecular basis of natural variation in light response, and suggest that some changes in light response in Arabidopsis and its relatives is adaptive. | ||
| Line 40: | Line 39: | ||
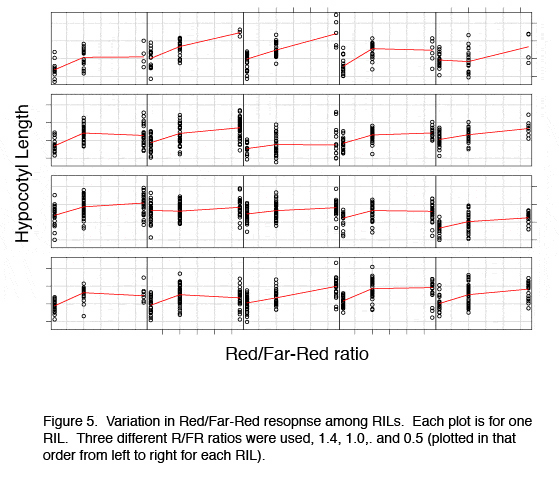
[[Image:Research_fig_5.jpg | center]] | [[Image:Research_fig_5.jpg | center | Figure 5]] | ||
In each environment we are measuring a variety of shade avoidance characters, including hypocotyl, stem, and petiole elongation, leaf angle, leaf shape, and flowering time. This work is part of an NSF funded collaboration with Cynthia Weinig at the University of Minnesota who is examining these lines under low and high competitive conditions in the field to study group versus individual selection. | In each environment we are measuring a variety of shade avoidance characters, including hypocotyl, stem, and petiole elongation, leaf angle, leaf shape, and flowering time. This work is part of an NSF funded collaboration with Cynthia Weinig at the University of Minnesota who is examining these lines under low and high competitive conditions in the field to study group versus individual selection. | ||
| Line 57: | Line 56: | ||
Recently developed likelihood models of codon based nucleotide substitution greatly increase the ability to detect positive selection and allow prediction of particular codons that have been subject to positive selection<sup>16-18</sup>. Thus, evolutionary data can be used to predict functionally interesting amino acid residues. PHYB is an ideal candidate for the application of these methods. While selective pressure on R/FR sensitivity could affect any gene in the PHYB pathway, there is evidence that PHYB itself is under selection. First, phys are evolving more rapidly than average plant genes<sup>19</sup>; second, we found that PHYB co-localizes with a quantitative trait locus (QTL) affecting response to white and red light in Arabidopsis<sup>10</sup>; and last, our unpublished analysis of ''A. thaliana'' and ''A. lyrata'' PHYB sequences suggests positive selection using the McDonald-Krietman test<sup>20</sup>. Codon based substitution models will be used to analyze PHYB sequence across the brassicaceae. Site-directed mutagenesis of interesting residues will be used to test the functional consequence of amino acid substitution using transgenic Arabidopsis and in vitro tests. This work is part of a HFSP funded collaboration with Ulrich Genick and Christian Fankhauser to explore variation in Phy structure and function. | Recently developed likelihood models of codon based nucleotide substitution greatly increase the ability to detect positive selection and allow prediction of particular codons that have been subject to positive selection<sup>16-18</sup>. Thus, evolutionary data can be used to predict functionally interesting amino acid residues. PHYB is an ideal candidate for the application of these methods. While selective pressure on R/FR sensitivity could affect any gene in the PHYB pathway, there is evidence that PHYB itself is under selection. First, phys are evolving more rapidly than average plant genes<sup>19</sup>; second, we found that PHYB co-localizes with a quantitative trait locus (QTL) affecting response to white and red light in Arabidopsis<sup>10</sup>; and last, our unpublished analysis of ''A. thaliana'' and ''A. lyrata'' PHYB sequences suggests positive selection using the McDonald-Krietman test<sup>20</sup>. Codon based substitution models will be used to analyze PHYB sequence across the brassicaceae. Site-directed mutagenesis of interesting residues will be used to test the functional consequence of amino acid substitution using transgenic Arabidopsis and in vitro tests. This work is part of a HFSP funded collaboration with Ulrich Genick and Christian Fankhauser to explore variation in Phy structure and function. | ||
==Bibliography== | ==Bibliography== | ||
Revision as of 18:03, 19 November 2006
Back to Maloof Lab