Altman:Research: Difference between revisions
David Altman (talk | contribs) No edit summary |
David Altman (talk | contribs) |
||
| (62 intermediate revisions by the same user not shown) | |||
| Line 3: | Line 3: | ||
<div style="padding: 10px; width: 700px; border: 5px solid #B22222;"> | <div style="padding: 10px; width: 700px; border: 5px solid #B22222;"> | ||
= | = Microscopic mosh pits: The role of force in regulating molecular motors = | ||
The inside of a cell is highly organized, and important players in a cell’s ability to maintain its ordered state are motor proteins. These microscopic engines allow a cell to transport, compartmentalize, and arrange its components by generating force and creating motion. Often, motors are envisioned as plodding along their filamentous protein track through a vast, empty cytoplasm of the cell. For example, we see this in the following gif of the motor protein kinesin moving along its microtubule track (from https://xvivo.com/examples/the-inner-life-of-the-cell/). | |||
[[File:KinesinWalking.gif|center|200px]] | |||
However, cells are remarkably dense and crowded, and are roiling with motion as thermal forces knock everything around. This becomes apparent, for example, in the following model of the bacterial cytoplasm, including the 50 most abundant types of macromolecules at experimentally measured concentrations (from McGuffee and Elcock. PLoS computational biology 6.3 (2010): e1000694.): | |||
<gallery position="center" widths=" | |||
[[File:journal.pcbi.1000694.s014_AMFvkY.gif|center|200px]] | |||
We thus can envision motors as generating their motion in the midst of a microscopic mosh pit. This is depicted nicely in the art of David Goodsell, who generated the following depiction (from https://www.interaliamag.org/articles/david-goodsell/ | |||
) of the transport of a vesicle filled with antibodies (yellow) by kinesin motors (extending diagonally from the vesicle). | |||
[[File:GoodsellKinesin.jpg|center|170px]] | |||
In the Altman lab, we study the myosin family of motor proteins, which move along the filamentous protein actin. | |||
[[File:Myosincartoonv2.jpg|center|300px]] | |||
We seek to understand how myosins are regulated by forces they experience in the crowded mosh pit of the cell. This work spans scales of complexity, from studies of individual purified motors to studies of ensembles of motors in cells. | |||
== Single molecule studies of myosins == | |||
We are examining the force-sensitivity of ''Acanthamoeba myosin 1c'' (AM1C) activity. Class 1 myosins have been split into two subclasses. While subclass 2 myosins are hypothesized to have a force-dependent activity, subclass 1 myosins are not. Two subclass 2 myosins ([http://www.ncbi.nlm.nih.gov/pubmed/18599791 Rat Myo1b] and [http://www.ncbi.nlm.nih.gov/pubmed/22908250 Mouse Myo1c]) have been shown to have a high degree of force-sensitivity, but no subclass 1 myosin has yet been tested. To test the force sensitivity of AM1C, which is from subclass 1, we are using an optical trap, which allows us to apply picoNewton forces to single myosin motors. | |||
<gallery position="center" widths="200px" heights="150px" > | |||
Image:opticaltrap.jpg|Cartoon of a myosin attached to a bead that is held in an optical trap. | Image:opticaltrap.jpg|Cartoon of a myosin attached to a bead that is held in an optical trap. | ||
Image:myosin1subclasses.jpg|Image taken from [http://www.ncbi.nlm.nih.gov/pubmed/15037306 De La Cruz, et al.] | Image:myosin1subclasses.jpg|Image taken from [http://www.ncbi.nlm.nih.gov/pubmed/15037306 De La Cruz, et al.] | ||
</gallery> | </gallery> | ||
We are conducting similar studies of myosin 6 with a mutation associated with hypertrophic cardiomyopathy (HCM) ([http://www.ncbi.nlm.nih.gov/pubmed/15060111 Myosin 6 H246R]). We seek to determine how the H246R mutation alters the chemomechanical cycle of the motor. | |||
<gallery position="center" widths="400px" heights="150px" > | |||
Image:Myosin6structure.tif|Crystal structure of the myosin 6 motor domain and insert-2 in the pre-powerstroke state. Insert-1 and insert-2 are shown in yellow and pink, respectively, and L310 and H246 are shown as spheres that are blue and green, respectively. In addition, bound MG-ADP-AlF4 is shown as a stick model. | |||
</gallery> | |||
== Retinal pigment epithelium phagocytosis == | == Retinal pigment epithelium phagocytosis == | ||
| Line 19: | Line 45: | ||
Retinal pigment epithelium (RPE) cells phagocytose waste shed by rod photoreceptor cells. This is important for the health of the eye, and failure to complete this process results in retinal degeneration. We study the role of molecular motors in this process. Specifically, we are testing the hypothesis that motor proteins myosin VI and VIIa generate the forces and motion required for the internalization of rod cell debris. To do this, we observe internalization of micron-sized microspheres by a primary RPE cell line (ARPE-19). To perturb the function of a particular myosin, we over-express myosins lacking the motor domain. | Retinal pigment epithelium (RPE) cells phagocytose waste shed by rod photoreceptor cells. This is important for the health of the eye, and failure to complete this process results in retinal degeneration. We study the role of molecular motors in this process. Specifically, we are testing the hypothesis that motor proteins myosin VI and VIIa generate the forces and motion required for the internalization of rod cell debris. To do this, we observe internalization of micron-sized microspheres by a primary RPE cell line (ARPE-19). To perturb the function of a particular myosin, we over-express myosins lacking the motor domain. | ||
<gallery position="center" widths="250px" heights="200px" > | |||
<gallery position="center" widths=" | |||
Image:RPEcell.jpg|Overlay of a DIC and fluorescence image of an ARPE-19 cell internalizing microspheres while transiently over-expressing a GFP-tagged myosin | Image:RPEcell.jpg|Overlay of a DIC and fluorescence image of an ARPE-19 cell internalizing microspheres while transiently over-expressing a GFP-tagged myosin | ||
Image:tracking_bead.jpg|Tracking a 1-micron-diameter microsphere that has been internalized by an ARPE-19 cell. | Image:tracking_bead.jpg|Tracking a 1-micron-diameter microsphere that has been internalized by an ARPE-19 cell. | ||
</gallery> | |||
== Techniques used in the lab == | |||
<gallery position="center" widths="250px" heights="200px" > | |||
Image:Motility.gif|Example of motility of fluorescent actin filaments driven by the myosin VI motors on the experimental surface. Total duration of movie is 10 minutes. | |||
Image:bead.gif|7.5 minute movie of a fluorescent bead in an ARPE-19 cell that is being trafficked toward the nucleus. Scale bar is 1 um. | |||
</gallery> | </gallery> | ||
</div> | </div> | ||
Latest revision as of 17:45, 4 February 2026
Microscopic mosh pits: The role of force in regulating molecular motors
The inside of a cell is highly organized, and important players in a cell’s ability to maintain its ordered state are motor proteins. These microscopic engines allow a cell to transport, compartmentalize, and arrange its components by generating force and creating motion. Often, motors are envisioned as plodding along their filamentous protein track through a vast, empty cytoplasm of the cell. For example, we see this in the following gif of the motor protein kinesin moving along its microtubule track (from https://xvivo.com/examples/the-inner-life-of-the-cell/).
However, cells are remarkably dense and crowded, and are roiling with motion as thermal forces knock everything around. This becomes apparent, for example, in the following model of the bacterial cytoplasm, including the 50 most abundant types of macromolecules at experimentally measured concentrations (from McGuffee and Elcock. PLoS computational biology 6.3 (2010): e1000694.):

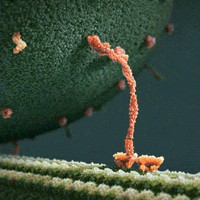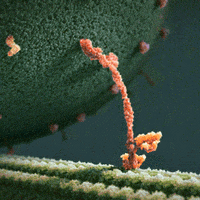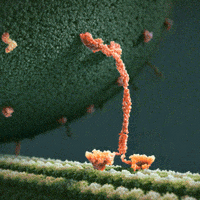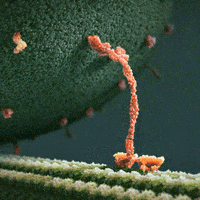
We thus can envision motors as generating their motion in the midst of a microscopic mosh pit. This is depicted nicely in the art of David Goodsell, who generated the following depiction (from https://www.interaliamag.org/articles/david-goodsell/
) of the transport of a vesicle filled with antibodies (yellow) by kinesin motors (extending diagonally from the vesicle).
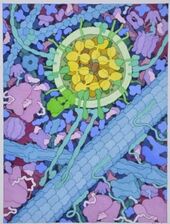
In the Altman lab, we study the myosin family of motor proteins, which move along the filamentous protein actin.
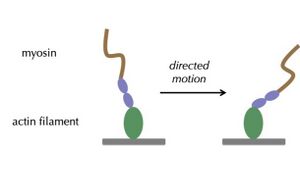
We seek to understand how myosins are regulated by forces they experience in the crowded mosh pit of the cell. This work spans scales of complexity, from studies of individual purified motors to studies of ensembles of motors in cells.
Single molecule studies of myosins
We are examining the force-sensitivity of Acanthamoeba myosin 1c (AM1C) activity. Class 1 myosins have been split into two subclasses. While subclass 2 myosins are hypothesized to have a force-dependent activity, subclass 1 myosins are not. Two subclass 2 myosins (Rat Myo1b and Mouse Myo1c) have been shown to have a high degree of force-sensitivity, but no subclass 1 myosin has yet been tested. To test the force sensitivity of AM1C, which is from subclass 1, we are using an optical trap, which allows us to apply picoNewton forces to single myosin motors.
-
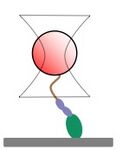
Cartoon of a myosin attached to a bead that is held in an optical trap.
-
Image taken from De La Cruz, et al.
We are conducting similar studies of myosin 6 with a mutation associated with hypertrophic cardiomyopathy (HCM) (Myosin 6 H246R). We seek to determine how the H246R mutation alters the chemomechanical cycle of the motor.
-
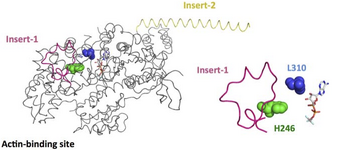
Crystal structure of the myosin 6 motor domain and insert-2 in the pre-powerstroke state. Insert-1 and insert-2 are shown in yellow and pink, respectively, and L310 and H246 are shown as spheres that are blue and green, respectively. In addition, bound MG-ADP-AlF4 is shown as a stick model.
Retinal pigment epithelium phagocytosis
Retinal pigment epithelium (RPE) cells phagocytose waste shed by rod photoreceptor cells. This is important for the health of the eye, and failure to complete this process results in retinal degeneration. We study the role of molecular motors in this process. Specifically, we are testing the hypothesis that motor proteins myosin VI and VIIa generate the forces and motion required for the internalization of rod cell debris. To do this, we observe internalization of micron-sized microspheres by a primary RPE cell line (ARPE-19). To perturb the function of a particular myosin, we over-express myosins lacking the motor domain.
-
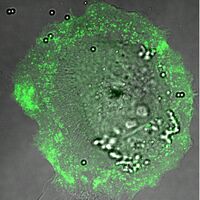
Overlay of a DIC and fluorescence image of an ARPE-19 cell internalizing microspheres while transiently over-expressing a GFP-tagged myosin
-
Tracking a 1-micron-diameter microsphere that has been internalized by an ARPE-19 cell.
Techniques used in the lab
-
Example of motility of fluorescent actin filaments driven by the myosin VI motors on the experimental surface. Total duration of movie is 10 minutes.
-
7.5 minute movie of a fluorescent bead in an ARPE-19 cell that is being trafficked toward the nucleus. Scale bar is 1 um.