Streptomyces:Research: Difference between revisions
Streptomyces (talk | contribs) No edit summary |
Streptomyces (talk | contribs) mNo edit summary |
||
| Line 2: | Line 2: | ||
<!--INSERT THE TEMPLATE HEADER--> | <!--INSERT THE TEMPLATE HEADER--> | ||
{{Streptomyces}} | {{Streptomyces}} | ||
{|cellspacing="5" cellpadding="10" style="background:#000; width: 800px;" | {|cellspacing="5" cellpadding="10" style="background:#000; width: 800px;" | ||
|-valign="top" | |-valign="top" | ||
| Line 22: | Line 21: | ||
*Cytoskeletal proteins | *Cytoskeletal proteins | ||
*Chromosome organisation during hyphal growth | *Chromosome organisation during hyphal growth | ||
<br> | <br/> | ||
---- | ---- | ||
| Line 31: | Line 30: | ||
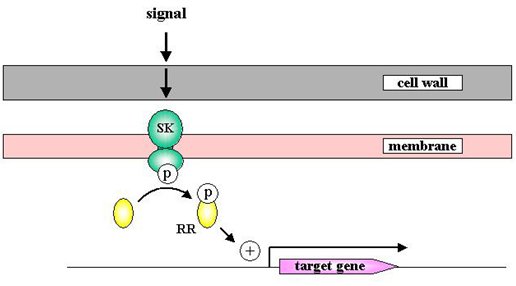
In order to survive, bacteria must sense and respond to their environment. One of the main ways in which bacteria do this is via two-component signal transduction pathways. In a typical two-component system the extracellular loop of the transmembrane sensor kinase senses a specific signal, autophosphorylates and passes that phosphate group to its cognate response regulator. The activated response regulator then switches on target genes to bring about a response to the original signal ('''Figure 1'''). | In order to survive, bacteria must sense and respond to their environment. One of the main ways in which bacteria do this is via two-component signal transduction pathways. In a typical two-component system the extracellular loop of the transmembrane sensor kinase senses a specific signal, autophosphorylates and passes that phosphate group to its cognate response regulator. The activated response regulator then switches on target genes to bring about a response to the original signal ('''Figure 1'''). | ||
<br> | <br/> | ||
<center> | <center> | ||
[[Image:Streptomyces_Matt_Figure1.jpg|frame|none|Figure 1. Classical model for a bacterial two-component signal transduction pathway.]] | [[Image:Streptomyces_Matt_Figure1.jpg|frame|none|Figure 1. Classical model for a bacterial two-component signal transduction pathway.]] | ||
</center> | </center> | ||
<br> | <br/> | ||
We have been studying these signal transduction pathways in the filamentous bacterium Streptomyces coelicolor. S. coelicolor is the model organism for the genus Streptomyces, a group of bacteria that produce about 80% of commercially important antibiotics. The streptomycetes are soil bacteria and undergo a complex life cycle that includes hyphal growth and sporulation. The S. coelicolor genome encodes 164 two-component proteins, more than nearly any other bacterium. This reflects both its complex life cycle and the highly variable soil environment in which it lives. We are investigating the signal transduction systems of S. coelicolor to work out how streptomycetes sense stresses (such as nutrient starvation and antibiotic attack from competing microorganisms) that trigger antibiotic production and sporulation. | We have been studying these signal transduction pathways in the filamentous bacterium Streptomyces coelicolor. S. coelicolor is the model organism for the genus Streptomyces, a group of bacteria that produce about 80% of commercially important antibiotics. The streptomycetes are soil bacteria and undergo a complex life cycle that includes hyphal growth and sporulation. The S. coelicolor genome encodes 164 two-component proteins, more than nearly any other bacterium. This reflects both its complex life cycle and the highly variable soil environment in which it lives. We are investigating the signal transduction systems of S. coelicolor to work out how streptomycetes sense stresses (such as nutrient starvation and antibiotic attack from competing microorganisms) that trigger antibiotic production and sporulation. | ||
<br> | <br/> | ||
<center> | <center> | ||
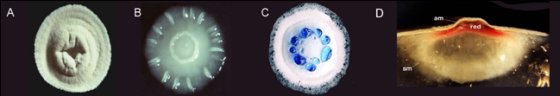
[[Image:Streptomyces_Matt_Figure2.jpg|frame|none|Figure 2. (A) wild type S. coelicolor, (B), a mutant which has lost the ability to sporulate, (C) a strain which overproduces the blue pigmented antibiotic actinorhodin and (D) a cross section of a colony showing the substrate mycelium (sm), aerial mycelium (am) and production of the antibiotic undecylprodigiosin, also known as “red”. Prodigiosins have recently been shown to have strong anti-cancer properties.]] | [[Image:Streptomyces_Matt_Figure2.jpg|frame|none|Figure 2. (A) wild type S. coelicolor, (B), a mutant which has lost the ability to sporulate, (C) a strain which overproduces the blue pigmented antibiotic actinorhodin and (D) a cross section of a colony showing the substrate mycelium (sm), aerial mycelium (am) and production of the antibiotic undecylprodigiosin, also known as “red”. Prodigiosins have recently been shown to have strong anti-cancer properties.]] | ||
</center> | </center> | ||
<br> | <br/> | ||
---- | ---- | ||
| Line 58: | Line 57: | ||
* DNA repair pathways of Ferroplasma acidarmanus - this archaea grows at 40 °C in the most acidic environment on earth (pH <1)! | * DNA repair pathways of Ferroplasma acidarmanus - this archaea grows at 40 °C in the most acidic environment on earth (pH <1)! | ||
* Relationship between DNA metabolism and genetic instability of DNA repeats | * Relationship between DNA metabolism and genetic instability of DNA repeats | ||
<br> | <br/> | ||
---- | ---- | ||
| Line 77: | Line 76: | ||
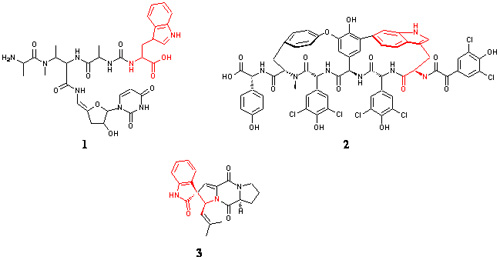
[[Image:Streptomyces_Rebecca_Figure1.jpg|frame|none| Figures 1, 2 & 3.]] | [[Image:Streptomyces_Rebecca_Figure1.jpg|frame|none| Figures 1, 2 & 3.]] | ||
</center> | </center> | ||
<br> | <br/> | ||
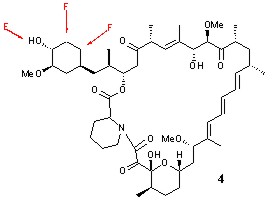
* Generation of novel fluorinated rapamycin analogues via a directed biosynthetic strategy | * Generation of novel fluorinated rapamycin analogues via a directed biosynthetic strategy | ||
Revision as of 11:18, 13 August 2007
Our research outline
<html> <script type="text/javascript" language="javascript"> var sc_project=2419278; var sc_invisible=1; var sc_partition=22; var sc_security="abf914b3"; </script> <script type="text/javascript" language="javascript" src="http://www.statcounter.com/counter/counter.js"></script><noscript><a href="http://www.statcounter.com/" target="_blank"><img src="http://c23.statcounter.com/counter.php?sc_project=2419278&java=0&security=abf914b3&invisible=0" alt="web metrics" border="0"></a> </noscript> <a href=http://www.theresonly1.eu> </a> <a href=http://www.theresonly1.co.uk> </a> <a href=http://www.rahart.eu> </a> <a href=http://www.jimandsarah.plus.com> </a> <a href=http://www.uea.ac.uk/~b101> </a> <a href=http://www.uea.ac.uk/~tdc07rfu> </a> </html>