SBB11Ntbk-Amy Li: Difference between revisions
No edit summary |
|||
| Line 16: | Line 16: | ||
==[[User:AMY Li|AMY Li]] 15:50, 8 March 2011 (EST)== | ==[[User:AMY Li|AMY Li]] 15:50, 8 March 2011 (EST)== | ||
[[Image:AmyGelPic3.JPG|thumb|right|A 1kb ladder was used, and the three parts have digestion sizes: (1) sbb1125 {P_spy} at 3744=613+3131bp, (2) sbb1133 {P_ycgE} at 3484=353+3131, and (3) sbb1138 {P_yciW} at 4619=1488+3131. Arrows denote which products were sequenced]] | |||
My '''colonies were picked''' over the weekend. | My '''colonies were picked''' over the weekend. | ||
| Line 36: | Line 38: | ||
Today, I '''ligated''' my three inserts with their corresponding vectors. <br> | Today, I '''ligated''' my three inserts with their corresponding vectors. <br> | ||
* Followed protocol for Ligation of EcoRI/BamHI digests: [[Template:SBB-Protocols Enz4|Ligation of EcoRI/BamHI digests]] <br> | |||
Then, I '''transformed''' E. coli cells with the plasmid product from ligation. <br> | Then, I '''transformed''' E. coli cells with the plasmid product from ligation. <br> | ||
* Followed protocol for Transformation by heat-shock: [[Template:SBB-Protocols Micro1|Transformation by heat-shock]] <br> | |||
* Letting shake in the 37 degree incubator for 45 minutes is sufficient (30 minutes bare minimum) <br> | |||
* Plated 100 ul of cell-plasmid cocktail on kanamycin antibiotics plate. <br> | |||
<br> | <br> | ||
| Line 47: | Line 49: | ||
Today, I '''redid my digestion''' because my digestion products mysteriously disappeared. <br> | Today, I '''redid my digestion''' because my digestion products mysteriously disappeared. <br> | ||
* Used 2 ul of 10x Dye (ran 10ul of stuff) <br> | |||
* Part sbb1133 (300+ bp) has a smeared band because of low concentration and may not work <br> | |||
<br> | <br> | ||
| Line 56: | Line 58: | ||
Today, I '''completed the digestion''' of my three PCR parts <br> | Today, I '''completed the digestion''' of my three PCR parts <br> | ||
* Followed protocol for Zymo Gel Purification: [[Template:SBB-Protocols Zymo3|Zymo Gel Purification]] <br> | |||
* Eluted PCR product in 8ul ddH2O | |||
<br> | <br> | ||
Next step: <br> | Next step: <br> | ||
* Ligation <br> | |||
<br> | <br> | ||
| Line 67: | Line 69: | ||
Today, I '''began the digestion''' of my three PCR products <br> | Today, I '''began the digestion''' of my three PCR products <br> | ||
* Performed the protocol for EcoRI/BamHI digestion of PCR Products: [[Template:SBB-Protocols Enz2|EcoRI/BamHI Digest of PCR Products]] <br> | |||
* Followed up with gel extraction with 1ul of 10x loading dye <br> | |||
* Stopped after melting gel in 600 ul ADB buffer | |||
<br> | <br> | ||
Next step: <br> | Next step: <br> | ||
* Continue gel extraction/purification to complete digestion and get pure DNA | |||
<br style="clear: both" /> | <br style="clear: both" /> | ||
Revision as of 19:28, 10 March 2011
AMY Li Quick Links
Amy Li's Project Page
Amy Li's Personal Page
BioE 140L Synthetic Biology Home Page
- The Synthetic Biology Protocols Page
- Spring 2011 BioE 140L Class Gel Pictures
~~!~~
AMY Li 14:06, 10 March 2011 (EST)
Today, I sent in two of each of my three products (six products total) for sequencing.
- All 50ul of purified ligation products were used
- Primers flank the gene and adhere onto the backbone
AMY Li 15:50, 8 March 2011 (EST)
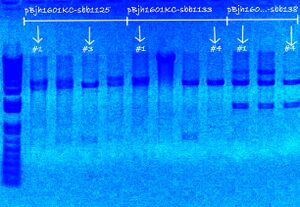
My colonies were picked over the weekend.
- Followed protocol for picking colonies: Picking of colonies
- Picked four colonies for each of my three plates
Today, I mini-prepped my colonies.
- Followed protocol for mini-prepping DNA: Miniprep purification of DNA
- Mini-prepped 12 culture tubes in total
I also digested my mini-preps.
- I did this to preliminarily check if my part sizes were correct before I send two of each part out for sequencing
- I threw out one darker-colored culture (remnants of a red bacteria suspected)
- This yields 11 mini-prep digestions and one ladder, filling up 12 wells in a gel
AMY Li 13:26, 3 March 2011 (EST)
Today, I ligated my three inserts with their corresponding vectors.
- Followed protocol for Ligation of EcoRI/BamHI digests: Ligation of EcoRI/BamHI digests
Then, I transformed E. coli cells with the plasmid product from ligation.
- Followed protocol for Transformation by heat-shock: Transformation by heat-shock
- Letting shake in the 37 degree incubator for 45 minutes is sufficient (30 minutes bare minimum)
- Plated 100 ul of cell-plasmid cocktail on kanamycin antibiotics plate.
AMY Li 15:31, 1 March 2011 (EST)
Today, I redid my digestion because my digestion products mysteriously disappeared.
- Used 2 ul of 10x Dye (ran 10ul of stuff)
- Part sbb1133 (300+ bp) has a smeared band because of low concentration and may not work
AMY Li 13:45, 24 February 2011 (EST)
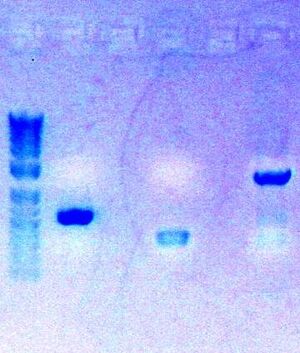
Today, I completed the digestion of my three PCR parts
- Followed protocol for Zymo Gel Purification: Zymo Gel Purification
- Eluted PCR product in 8ul ddH2O
Next step:
- Ligation
AMY Li 13:30, 22 February 2011 (EST)
Today, I began the digestion of my three PCR products
- Performed the protocol for EcoRI/BamHI digestion of PCR Products: EcoRI/BamHI Digest of PCR Products
- Followed up with gel extraction with 1ul of 10x loading dye
- Stopped after melting gel in 600 ul ADB buffer
Next step:
- Continue gel extraction/purification to complete digestion and get pure DNA
AMY Li 13:29, 17 February 2011 (EST)
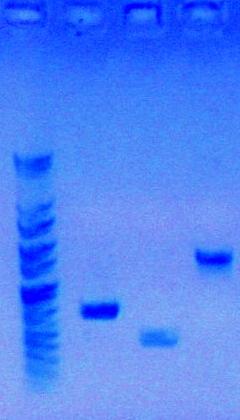
This is the roadmap for my project:
For each of my three parts:
0) Amplify part using PCR
1) Analytical Gel to see if my PCR products were successful
2) Zymo Clean-up to isolate only DNA part
3) Digestion using EcorRI and BamHI
4) Ligation
5) Transformation
Today, I retrieved my three PCR products and ran them through analytical gel.
- Ran an analytical gel on my PCR products to visualize them
- Used 5ul Loading Dye and 2ul PCR product
- Not using Preparation Gel because don't need to gel extract
Results of Analytical Gel: PCR products were successful!
- Well 1: PCR Part sbb1125 {P_spy} 629 bp
- Well 2: PCR Part sbb1138 {P_yciW} 1504 bp
- Well 3: PCR Part sbb1133 {P_ycgE} 369 bp
Today I also column purified my PCR products by Zymo Clean-up
- Followed protocol for Regular Zymo Clean-up: Regular Zymo Cleanup
- For all "spin through"s, spun at 30 seconds at 14.5rpm (max RPM)
- Eluted each part with 33ul ddH20
Next Steps:
Digestion with EcorRI/BamHI, Ligation, then Transformation.
AMY Li 17:04, 15 February 2011 (EST)
Today, I set up three cloning by PCR reactions, one for each of my three promoter parts.
- Followed the protocol for cloning DNA: Cloning by PCR
- Resuspended ALI_005 to 100uM and diluted it to 10uM
- Used the Expand buffer and polymerase for my PCR because the part templates were all from MG1655 gDNA
- PCR Thermocycler program 2K55 because all PCR products are less than 2kb
AMY Li 17:04, 15 February 2011 (EST)
Construction Files for my three Bgl Bricks basic parts for the random ball project:
Part sbb1125 {P_spy}
PCR ss51r/ss51f on MG1655 gen. (629bp, EcoRI/BamHI)
Sub into pBjh1601KC-Bca1144#5 (EcoRI/BamHI, 3131+910, L)
Product is pBjh1601KC-sbb1125 {P_spy}
ss51f Reverse Cloning of P_spy tttggGGATCCcatgtcctgatgcggaccgaacttgcc
ss51r Forward Cloning of P_spy aaaccGAATTCatgAGATCTtggcgcaggacggagaggaacg
Part sbb1138 {P_yciW}
PCR ss65r/ss65f on MG1655 gen. (1504bp, EcoRI/BamHI)
Sub into pBjh1601KC-Bca1144#5 (EcoRI/BamHI, 3131+910, L)
Product is pBjh1601KC-sbb1138 {P_yciW}
ss65f Reverse Cloning of P_yciW tttggGGATCCgtagaaagggatcgctgacc
ss65r Forward Cloning of P_yciW aaaccGAATTCatgAGATCTcggtgctggagaatattctgcagg
Part sbb1133 {P_ycgE}
PCR ss60f/ALI006 on E. coli MG1655 gDNA (369 bp, EcoRI/BamHI)
Sub into pBjh1601KC-Bca1144#5 (EcoRI/BamHI, 910+3131 bp, L)
Product is pBjh1601KC-sbb1133 {P_ycgE}
ALI005 Forward cloning of P_ycgE CCATAgaattcatgagatctGTTTGCTAAAGCTAAATTGAATGGTATCC
ss60f Reverse cloning of P_ycgE tttggGGATCCggcgttgccaggcccggagagtgacag
AMY Li 23:04, 7 February 2011 (EST)
Hello World!