BioMicroCenter:Genome Seq: Difference between revisions
| Line 3: | Line 3: | ||
== Genome Seq == | == Genome Seq == | ||
[[Image:BioMicroCenter-72_72_errorRate.png | 400px | right]] | [[Image:BioMicroCenter-72_72_errorRate.png | 400px | right]] | ||
Both the HiSeq and the Genome Analyzer support '' | Both the HiSeq and the Genome Analyzer support ''de novo'' sequencing with extended read lengths, high passed-filter read counts, and the option to run lanes as paired-end for additional read length and long-range sequence information. | ||
== Sample Submission Guidelines == | == Sample Submission Guidelines == | ||
Revision as of 13:55, 30 September 2011
| HOME -- | SEQUENCING -- | LIBRARY PREP -- | HIGH-THROUGHPUT -- | COMPUTING -- | DATA MANAGEMENT -- | OTHER TECHNOLOGY |
Genome Seq
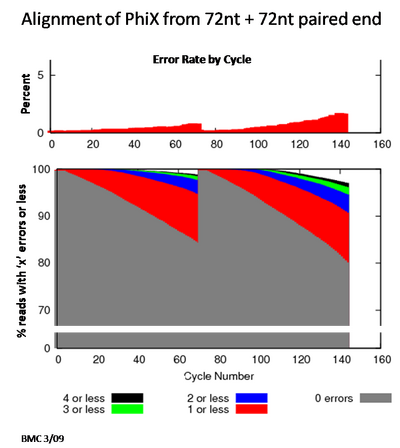
Both the HiSeq and the Genome Analyzer support de novo sequencing with extended read lengths, high passed-filter read counts, and the option to run lanes as paired-end for additional read length and long-range sequence information.
Sample Submission Guidelines
Users may submit unprepared libraries to the BioMicro Center for Sample Prep followed by Illumina sequencing, or they may submit prepared libraries directly to sequencing. Standard Illumina protocols for sample preparation can be found here. Illumina Paired End sample preparation kits can be purchased directly from Illumina or through the BioMicro Center for $3,600.00 plus shipping costs (10 sample preps per kit). To request an order or for more information please contact Ali Perrotta
Samples which are ready for sequencing should be gel-purified according to your protocol, and provided as at least 5uL of solution at a concentration of at least 10 nM in EB/ Tween 0.1%. Quality control will be run on all submitted samples.
Samples generated using standard kits with Illumina PCR primers can be submitted without sequencing primers. Custom sample preparations using customized PCR primer sequences should be provided along with a complementary sequencing primer.
Data
Images acquired from the solexa sequencer are processed through the bundled solexa inage extraction pipeline to get the sequence and quality score for each base. The data is aligned to a reference genome, if requested, using an interative ELAND algorithm. An in-depth QC report is included in the package. .
Genome Seq Literature
<biblio>
- Paper1 pmid=18677319
