Prince:Proteomics: Difference between revisions
From OpenWetWare
Jump to navigationJump to search
| Line 10: | Line 10: | ||
[[Prince:Filter-aided Sample Preparation|FASP (Filter-aided Sample Preparation)]] | [[Prince:Filter-aided Sample Preparation|FASP (Filter-aided Sample Preparation)]] | ||
[[Prince:Strong Cation Exchange| SCX (Strong Cation Exchange)]] | |||
[[Prince:Phospho-protein Enrichment|Phospho-protein Enrichment]] | [[Prince:Phospho-protein Enrichment|Phospho-protein Enrichment]] | ||
Revision as of 20:07, 3 February 2012
Introduction
- Proteomics is the study of protein structure and function. Mass spectrometry measures the m/z (mass to charge) ratio of particles. So mass spectrometry based proteomics is the study of protein structure and function by identifying the m/z ratio of proteins or pieces of proteins called peptides. Mass spectrometry can identify thousands of proteins in a single analysis. Our goal is to identify and quantitate changes in protein concentration and phosphorylation of proteins in the PAS kinase network as a result of PAS kinase perturbation.
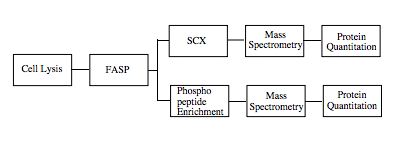
Experiment Design
FASP (Filter-aided Sample Preparation)