20.109: 2-component signaling
Module ideas
- Day 1: set up dark/light system in plates, liq culture
- Day 2: b-gal assay of cultures, set up photo
- Day 3: transform library and screen, recapitulate setup electronically
- Day 4: re-streak candidate, journal article discussion <teacher ONs in light and dark>
- Day 5: protein gel and blot (or RNA isolation) and b-gal assay while running
- Day 6: probe blot, also miniprep culture and send DNA for sequencing
- Day 7: Journal club, data back
following week written work due
Useful links
- Original module
- b-gal assay
- SDM kitSDM lab protocol
- controlled swimmingchemotaxis lab protocol
- APE
- phenotype arrays
- q-PCR
Useful Images and Files




References
- Synthetic biology: engineering Escherichia coli to see light. Levskaya A, Chevalier AA, Tabor JJ, Simpson ZB, Lavery LA, Levy M, Davidson EA, Scouras A, Ellington AD, Marcotte EM, Voigt CA.
Nature. 2005 Nov 24;438(7067):441-2. PMID: 16306980 and pdf
- Engineered single- and multi-cell chemotaxis pathways in E. coli. Goldberg SD, Derr P, DeGrado WF, Goulian M. Mol Syst Biol. 2009;5:283. Epub 2009 Jun 16. PMID: 19536206 and pdf
- Specificity in two-component signal transduction pathways. Laub MT, Goulian M. Annu Rev Genet. 2007;41:121-45. Review. PMID: 18076326 and pdf
- Mutations that alter the kinase and phosphatase activities of the two-component sensor EnvZ. Hsing W, Russo FD, Bernd KK, Silhavy TJ. J Bacteriol. 1998 Sep;180(17):4538-46. PMID: 9721293 and pdf
- Cross-talk suppression between the CpxA-CpxR and EnvZ-OmpR two-component systems in E. coli. Siryaporn A, Goulian M. Mol Microbiol. 2008 Oct;70(2):494-506. Epub 2008 Aug 29. PMID: 18761686 and pdf
- Design and signaling mechanism of light-regulated histidine kinases. Möglich A, Ayers RA, Moffat K. J Mol Biol. 2009 Feb 6;385(5):1433-44. Epub 2008 Dec 14. PMID: 19109976 and pdf
- EnvZ-OmpR interaction and osmoregulation in Escherichia coli. Cai SJ, Inouye M. J Biol Chem. 2002 Jul 5;277(27):24155-61. Epub 2002 Apr 24. PMID: 11973328 and pdf
K-P+ and K+P- mutations
06.16.09 email from Mike: As for the EnvZ mutants, here's the list we've compiled from the literature. Keep in mind that K+P- really means a shift in the balance of kinase and phosphatase activities and similarly for the K-P+ alleles. None of them is perfectly "clean" in eliminating one of the activities:
K+P-
- V241G = V555G in Cph8
- G240E = G554E in Cph8
- S242D = S556D in Cph8
- P248Q
- T247R
- Q283P
- Y287D
- L288P
K-P+
- A239T = A553 in Cph8
- N343K
- F390L
- H243A = H557A in Cph8
NK note: relevant reference:
- Mutations that alter the kinase and phosphatase activities of the two-component sensor EnvZ. Hsing W, Russo FD, Bernd KK, Silhavy TJ. J Bacteriol. 1998 Sep;180(17):4538-46. PMID: 9721293
- The role of the G2 box, a conserved motif in the histidine kinase superfamily, in modulating the function of EnvZ. Zhu Y, Inouye M. Mol Microbiol. 2002 Aug;45(3):653-63. PMID: 12139613


Primers
NO281 = KP randomized template
- 5' CATATGGCGGCTGGTGTTAAGCAACTGGCGGATGACCGCACGCTGCTGATG RNS RNS RNS RNS SNW GACTTGCGCACGCCGCTGACGCGT
NO284 = P+ randomized template
- 5' CATATGGCGGCTGGTGTTAAGCAACTGGCGGATGACCGCACGCTGCTGATG RNS GGG GTA AGT SNW GACTTGCGCACGCCGCTGACGCGT
NO285 = K+ randomized template
- 5' CATATGGCGGCTGGTGTTAAGCAACTGGCGGATGACCGCACGCTGCTGATG GCG RNS RNS RNS CAC GACTTGCGCACGCCGCTGACGCGT
NO282 = KPLibrary_fwd_1535_1573
- 5' CAGAAGAATTGCATATGGCGGCTGGTGTTAAGCAACTGG
- 39 mer
- Tm (all) = 66.3 ºC
- Tm (landing, 28 mer) = 63.3 ºC
NO283 = KPLibrary_rev_1643_1609
- 5' AGGCGAATACGCGTCAGCGGCGTGCGCAAGT
- 31 mer
- Tm (all) = 72.1 ºC
- Tm (landing, 23 mer) = 70.5 ºC
NO279 = GFPfiller_Nde_fwd
5' CATTAGCATATGGGATCCTAAGAGGGCGAGGGCGATGCCACC
- 42 mer
- Tm (all) = 69.7 ºC
- Tm (landing, 24 mer) = 67 ºC
NO280 = GFPfiller_Mlu_rev
5' CATTAGTGCGCAGATATCTTACTTGTACAGCTCGTCCATGC
corrected: 5' CATTAGACGCGTGATATCTTACTTGTACAGCTCGTCCATGC
- 41 mer
- Tm (all) = 64.6 ºC
- Tm (landing, 23 mer) = 57 ºC
NO288= GFPfiller_Nde_rev
5'CATTAGCATATGGATATCTTACTTGTACAGCTCGTCCATGC
- Tm (all) = 62.6 ºC
- Tm (landing, 23 mer) = 57 ºC
Possible ways to epitope tag
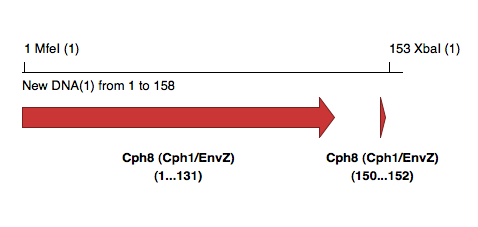 All tags at C-terminus, with intention to clone product using unique MfeI and XbaI in pCph8
All tags at C-terminus, with intention to clone product using unique MfeI and XbaI in pCph8
HA
- 9-amino acid sequence (YPYDVPDYA) recognized by the anti-12CA5 (=anti-HA)
- reverse translate at |Gene Design with E. coli codon bias
- default settings gives sequence: TAC CCG TAC GAC GTT CCG GAC TAC GCT
caattgTGCAGCGTATCGTGGATAACCATAACGGGATGCTGGAGCTTGGCACCAGCGAGCGGGGCGGGCTTTCCATTCGCGCCTGGCTGCCAGTGCCGGTAACGCGGGCGCAGGGCATGACAAAAGAAGGGTACCCGTACGACGTTCCGGACTACGCTTAAtctaga
to PCR from DNA2.0 vector
- NO286=HAfrag_fwd
- 5' GTCGCTGAACAATTGTGCAGCGTATCGTGG
- Tm = 61.0 ºC
- NO287=HAfrag_rev
- 5' GAACTCGATTGACGTCTAGATTAAGCGTAG
- Tm = 58.1 ºC
to check insert of HA in pCph8
- NO289 = Cph8+HAseqprimer_bp135_rev
- 5' ATTACCGCCTTTGAGTGAGC
His6
- 6 Histidine sequence from pET-45b(+) bp 4963-4980 (Novagen product)
caattgTGCAGCGTATCGTGGATAACCATAACGGGATGCTGGAGCTTGGCACCAGCGAGCGGGGCGGGCTTTCCATTCGCGCCTGGCTGCCAGTGCCGGTAACGCGGGCGCAGGGCATGACAAAAGAAGGGcatcaccaccaccatcacTAAtctaga
