BME103:T130 Group 2: Difference between revisions
| Line 161: | Line 161: | ||
'''Some basic components involved in a PCR reaction are:'''<br> | '''Some basic components involved in a PCR reaction are:'''<br> | ||
'''Template DNA:''' the original strand of DNA that is going to be copied<br> | '''Template DNA:''' This DNA template is the original strand of DNA that is going to be copied<br> | ||
'''Primers:''' attach to the sites on the DNA strands that are at either end of the template DNA | '''Primers:''' These primers attach to the sites on the DNA strands that are at either end of the template DNA. They drive the PCR because they are powerful enzymes that allow replication to occur. In this reaction, if primers are able to attach to the DNA strand, the patient is negative for cancer.<br> | ||
'''Taq Polymerase:''' reads the DNA code and matches new nucleotides with the template strand | '''Taq Polymerase:''' Tag Polymerase reads the DNA code and matches new nucleotides with the template strand. It bind the pairs together with hydrogen bonds, for example A's are bonded to T's and C's are bonded to G's. These base pairs make it possible to make several copies of the DNA strands. <br> | ||
'''Magnesium Chloride:''' a three atom molecule that binds to Taq | '''Magnesium Chloride:''' Magnesium Chloride is a three atom molecule that binds to Taq Polymerase as a co-factor to help Tag Polymerase it function properly.<br> | ||
'''dNTP's:''' | '''dNTP's:''' Deoxinucleotide Triphosphates are the new nucleotides<br><br> | ||
'''A PCR functions by going through a series of thermal cycles:'''<br> | '''A PCR functions by going through a series of thermal cycles:'''<br> | ||
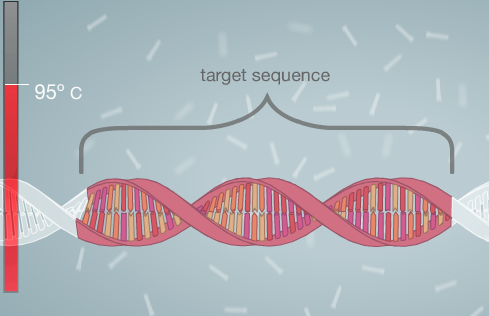
In the first cycle the PCR machine heats up to 95 degrees Celsius. This unzips the DNA strand creating two separate DNA strand molecules and exposes the nucleotides we wish to look at. In the second cycle the temperature drops to 57 degrees Celsius which allows the | In the first cycle, the PCR machine heats up to 95 degrees Celsius. This unzips the DNA strand creating two separate DNA strand molecules and exposes the nucleotides that we wish to look at. In the second cycle, the temperature drops to 57 degrees Celsius which allows the primers to bind to the ends of the DNA strands. The third cycle heats up the DNA to 72 degrees Celsius to allow DNA replication by the enzyme Taq Polymerase. These three cycles are repeated many more times, allowing for billions of DNA copies to be made. We need multiple copies of the DNA so that we can see if the patient tests positive for cancer.<br><br> | ||
'''r17879961'''<br> | '''r17879961'''<br> | ||
The single nucleotide polymorphism, or SNP, that codes for cancer is AACTCTTACA'''C'''TCGATACAT; in a normal patient, the C is replaced with T | The single nucleotide polymorphism, or SNP, that codes for cancer is AACTCTTACA'''C'''TCGATACAT; in a normal patient, the '''C''' is replaced with '''T'''(AACTCTTACA'''T'''TCGATACAT). The backwards part to these two sequences is TTGAGAATGT'''A'''AGCTATAGTA. The reason the C causes cancer in a patient is because there is an improper base pair causing the primers not to bind. This results in the formation of single strands. Keep in mind that C does not code wit A, it only codes with C. After the PCR's thermal cycles are finished the cancerous patient's DNA will be single stranded and we will be able to see this after SYBR green dye is added and the sample appears green. The SYBR green dye does not express the double stranded DNA, non-cancerous patients. <br><br> | ||
'''Reliability: Bayes Rule:'''<br> | '''Reliability: Bayes Rule:'''<br> | ||
We saw that the SNP is two hundred base pairs away, meaning every two hundred base pairs the missense mutation is present. <br> | We saw that the SNP is two hundred base pairs away, meaning every two hundred base pairs the missense mutation is present. <br> | ||
Revision as of 20:22, 14 November 2012
| Home People Lab Write-Up 1 Lab Write-Up 2 Lab Write-Up 3 Course Logistics For Instructors Photos Wiki Editing Help | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
OUR TEAMLAB 1 WRITE-UPInitial Machine TestingThe Polymerase Chain Reaction (PCR) machine is used to test for variations in nucleotides. It is simple, easy to use, and portable. For our purposes we used the PCR machine to test for cancer in patients. It was able to test up to sixteen test tubes at a time and took slightly more than an hour and a half to complete the cycles. It heats up in order to separate the DNA strand and then cools to allow primers to attach and new replicated strands to form. It then heats back up to repeat the same cycles.
Experimenting With the Connections When we unplugged the LCD screen from the PCR machine, the machine's display screen stopped working, but information continued to transmit to the computer. When we unplugged the white wire that connects the Thermocouple to the 16-tube PCR block, the machine's temperature measurement was disabled, in other words, it no longer kept track of heat readings.
On October 18th and 20th the PCR machine responded normally with no signs of malfunction. However, it took a little bit longer to complete the cycle; specifically, it took an hour and forty minutes in total. The machine LED display matched the computer screen which was a sign of success and proper function.
Protocols
Sample 1: Positive Control (contains cancer DNA template), Tube label: P
Using a Fluorimeter
Fluorimeter Assembly Procedures:
ImageJ Set Up Procedures:
Research and DevelopmentSpecific Cancer Marker Detection - The Underlying Technology
Some basic components involved in a PCR reaction are: r17879961
Results
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||