Biomod/2012/UTokyo/UT-Hongo/Assembly: Difference between revisions
No edit summary |
No edit summary |
||
| Line 2: | Line 2: | ||
= | In this section, we write about our experiment. | ||
=Assembly of the DNA Shell= | |||
First, we mixed scaffold (used base sequence of M13) and staples, and check whether DNA origami was hybridized as we had designed by agarose gel electrophoresis and atomic force microscope. | |||
Scaffold DNA and staples were in 1X TE buffer. The staple DNA was mixed with M13mp18 (160nM of each staple DNA, 1.6nM scaffold DNA, a 100-fold excess of staple DNA) in 1X TAE/Mg buffer and annealed from 95℃ to 20℃ in a themal cycler at a rate of 6.25℃ per 10minute 12 steps. | |||
==Agarose Gel Electrophoresis== | ==Agarose Gel Electrophoresis== | ||
' Samples were electrophoreses in a 0.6% agarose gel containing 1xTAE/Mg buffer. The agarose gel was run at 27℃. ' | '' Samples were electrophoreses in a 0.6% agarose gel containing 1xTAE/Mg buffer. The agarose gel was run at 27℃. '' | ||
<center> | <center> | ||
| Line 53: | Line 54: | ||
===Outlook=== | ===Outlook=== | ||
First picture shows that the origami with all staples was formed as designed. Also, these pictures may show that the first origami differ from the second one in the structure between two squares. | First picture shows that the origami with all staples was formed as designed. Also, these pictures may show that the first origami differ from the second one in the structure between two squares. | ||
=Capturing ability= | |||
Next, we ascertained whether DNA Shell ccaptured target molecules by agarose gel electrophoresis, fluorometry, and AFM. | |||
==Agarose gel electrophoresis== | |||
==Fluorometry== | |||
==AFM== | |||
=Supporting enzyme= | |||
Based on the results mentioned above, we did further advanced experiment. We ascertained whether DNA Shell supported enzymes. | |||
==Background== | |||
We used tetramethylbenzidine (TMB), streptavidin with horseradish peroxidase (HRP) labeling and trypsin. TMB can be oxidized for the reduction of hydrogen peroxide to water by peroxidase enzymes such as HRP. When TMB is oxidized, the color of the solution takes on a blue color. If pH of the solution is low (for example, using sulfric acid), the color turns into yellow. The former blue color can be read at a wavelength of nm and the latter yellow color can be read at nm. | |||
==Experiment condition== | |||
Revision as of 09:34, 15 October 2012
<html> <head> <style> <!-- HIDE WIKI STUFF -->
- column-one { display:none; width:0px;}
.container{background-color: #ffffff; margin-top:0px} .OWWNBcpCurrentDateFilled {display: none;}
- content { width: 0px; margin: 0 auto auto 0; padding: 0em 0em 0em 0em; align: center;}
- column-content {width: 0px; float: left; margin: 0 0 0 0;padding: 0;}
.firstHeading {display:none; width:0px;}
- globalWrapper{width:984px; background-color: #ffffff; margin-left: auto; margin-right: auto}
- column-one {display:none; width:0px;background-color: #ffffff;}
- content{ margin: 0 0 0 0; align: center; padding: 12px; width: 960px;background-color: #ffff; border: 0;}
- bodyContent{ width: 960px; align: center; background-color: #ffffff;}
- column-content{width: 984px;background-color: #ffff;}
- footer{display: none; position: center; width: 960px}
@media screen {
body { background: #ffffff url(http://openwetware.org/images/1/14/Biomod-2012-utokyo-uthongo-wrapper-bg.jpg) repeat; }
}
</style>
<style>
- {
text-shadow: 0px 1px 2px #aaa; }
- bodyContent, #column-content, #content {
background: #ffffff url(http://openwetware.org/images/1/14/Biomod-2012-utokyo-uthongo-wrapper-bg.jpg) repeat; }
- column-content {
border-left: 1px solid #888; border-right: 1px solid #888;
}
- header {
width: 960px; height: 304px; background: url(http://openwetware.org/images/3/3b/Biomod-2012-UTokyo-UT-Hongo_header_test2.png) no-repeat; background-size: 100% auto; margin-bottom: 0px; padding: 0px 0px 0px 0px;
/* border-bottom: 1px solid #DEDEDE; */ }
- menu {
display: block; width: 956px; padding: 0px; background-color: black; border: 2px solid #888; margin-bottom: 15px;
}
ul.menu li a {
display: block;
/* border: 1px solid #474655; */
padding: 8px 10px; text-decoration: none;
/* color: #333; */
color: #fff;
/* width: 121px; */
margin: 0px; text-align: center; font-size: large;
}
ul.menu li a:hover{
color: cyan;
}
ul.menu { /* display: none;
position: absolute; */ height: 40px; margin:0; padding:0; list-style:none; float: center;
}
ul.menu li { /* display:inline;*/
margin:0; padding:6px; float: left; position: relative;
}
ul.menu li:hover ul {
display: block; position: absolute; z-index: 100;
}
ul.submenu {
display: none; list-style: none; background-color: #f0f0f0; border: 1px solid #ccc;
}
ul.submenu li {
float: none; margin: 0px;
}
ul.submenu li a {
text-align: left; margin: 0px; width: 340px; padding: 0 0 0 12px; color: black; text-decoration: none; font-size: large
}
ul.submenu li a:hover {
color: black; text-decoration: underline;
}
.thumb {
border-image: url(http://openwetware.org/images/1/14/Biomod-2012-utokyo-uthongo-wrapper-bg.jpg) repeat;
}
.mytable {
border-image: url(http://openwetware.org/images/1/14/Biomod-2012-utokyo-uthongo-wrapper-bg.jpg) repeat;
}
</style> </head> <body>
<!--<div style="text-align: right"> <p><a href="http://openwetware.org/index.php?title=Template:Biomod/2012/UTokyo/UT-Hongo&action=edit">edit this header</a></p> </div>-->
<div style="height: 60px; margin-bottom: 18px;">
<div style="float: left; width: 250px; padding: 3px; background-color: white;">
<a href="http://biomod.net/"><img src="http://openwetware.org/images/8/82/Biomod2012-logo.png" width="250px" height="50px"></img></a>
</div>
<div style="float: right; width: 240px; padding: 3px; background-color: white;">
<a href="http://www.u-tokyo.ac.jp/en/" target="_blank"><img src="http://www.u-tokyo.ac.jp/en/images/banner/UT-logo.gif" width="234" height="60" border="0" alt="The University of Tokyo"></img></a>
</div>
</div>
<div id="header"> </div>
<div id="menu">
<ul class="menu">
<li class="toppage"><a href="/wiki/Biomod/2012/UTokyo/UT-Hongo">Top</a></li> <li class="motives"><a href="/wiki/Biomod/2012/UTokyo/UT-Hongo/Intro">Motives</a></li> <!-- <li class="design"><a href="/wiki/Biomod/2012/UTokyo/UT-Hongo/Function">Design</a></li> --> <li class="result"><a href="/wiki/Biomod/2012/UTokyo/UT-Hongo/Assembly">Design & Results</a> <ul class="submenu"> <li><a href="/wiki/Biomod/2012/UTokyo/UT-Hongo/Assembly#Assembly_of_the_DNA_Shell">Assembly of the DNA Shell</a></li> <li><a href="/wiki/Biomod/2012/UTokyo/UT-Hongo/Assembly#Capturing_ability">Capturing Ability</a></li> <li><a href="/wiki/Biomod/2012/UTokyo/UT-Hongo/Assembly#Immobilizing_on_microfluidic_device">Immobilizing on microfluidic device</a></li> <li><a href="/wiki/Biomod/2012/UTokyo/UT-Hongo/Assembly#Supporting_Enzyme">Supporting Enzyme</a></li> </ul> </li> <li class="method"><a href="/wiki/Biomod/2012/UTokyo/UT-Hongo/Method">Method</a></li> <li class="futurework"><a href="/wiki/Biomod/2012/UTokyo/UT-Hongo/FutureWork">Progress & Beyond</a></li> <li class="team"><a href="/wiki/Biomod/2012/UTokyo/UT-Hongo/Team">Team</a></li> <li class="acknowledgement"><a href="/wiki/Biomod/2012/UTokyo/UT-Hongo/Acknowledgement">Acknowledgement</a></li>
</ul> </div>
</body> </html>
In this section, we write about our experiment.
Assembly of the DNA Shell
First, we mixed scaffold (used base sequence of M13) and staples, and check whether DNA origami was hybridized as we had designed by agarose gel electrophoresis and atomic force microscope.
Scaffold DNA and staples were in 1X TE buffer. The staple DNA was mixed with M13mp18 (160nM of each staple DNA, 1.6nM scaffold DNA, a 100-fold excess of staple DNA) in 1X TAE/Mg buffer and annealed from 95℃ to 20℃ in a themal cycler at a rate of 6.25℃ per 10minute 12 steps.
Agarose Gel Electrophoresis
Samples were electrophoreses in a 0.6% agarose gel containing 1xTAE/Mg buffer. The agarose gel was run at 27℃.
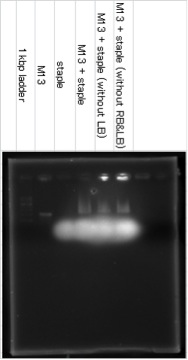
The band of M13+staple ran longer than the band of M13, it showed that the structure which had big molecular mass was created. The lower fuzzy band was the band of excess staple DNA. RB and LB means Right Bridge and Left Bridge. These connect three squares of DNA origami.
AFM
These DNA origami (M13 + all staples, M13 + all staples except ones which combine with M13 between the squares (except LB and RB), M13 + all staples except one which combine with M13 between the center square and the right square (except RB)) were observed by using atomic force microscope (AFM) to confirm that these DNA origami were formed correctly. 1xTAE/Mg solution was utilized as buffer.
M13+staple (represent 500nm at one side)



M13+staple (without LB,RB)



M13+staple (without RB)
Now trying..
Outlook
First picture shows that the origami with all staples was formed as designed. Also, these pictures may show that the first origami differ from the second one in the structure between two squares.
Capturing ability
Next, we ascertained whether DNA Shell ccaptured target molecules by agarose gel electrophoresis, fluorometry, and AFM.
Agarose gel electrophoresis
Fluorometry
AFM
Supporting enzyme
Based on the results mentioned above, we did further advanced experiment. We ascertained whether DNA Shell supported enzymes.
Background
We used tetramethylbenzidine (TMB), streptavidin with horseradish peroxidase (HRP) labeling and trypsin. TMB can be oxidized for the reduction of hydrogen peroxide to water by peroxidase enzymes such as HRP. When TMB is oxidized, the color of the solution takes on a blue color. If pH of the solution is low (for example, using sulfric acid), the color turns into yellow. The former blue color can be read at a wavelength of nm and the latter yellow color can be read at nm.