Brown BIOL1220:Notebook/SynBio in Theory and Practice/Genes/ADAR: Difference between revisions
From OpenWetWare
Jump to navigationJump to search
Elias Scheer (talk | contribs) |
Elias Scheer (talk | contribs) No edit summary |
||
| Line 2: | Line 2: | ||
=== The Functions of the ADAR Family === | === The Functions of the ADAR Family === | ||
* Three members of this deaminase family are known: ADAR 1, ADAR 2, and ADAR 3 that share a common modular domain structure. | * Three members of this deaminase family are known: ADAR 1, ADAR 2, and ADAR 3 that share a common modular domain structure. | ||
[[Image:ADAR_pic.tiff]] | |||
* ADARs modify pre-mRNA and recognize structural determinants in the RNA. | * ADARs modify pre-mRNA and recognize structural determinants in the RNA. | ||
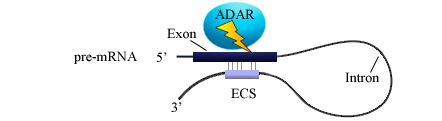
* In mice, the editosomes with ADAR proteins require some cis-acting elements like an intronic 'editing-site complementary sequence (ECS)'. Although evolutionarily conserved, the actual role of ECS is not yet elucidated in humans. ''A representation of the editing complex is shown below.'' | * In mice, the editosomes with ADAR proteins require some cis-acting elements like an intronic 'editing-site complementary sequence (ECS)'. Although evolutionarily conserved, the actual role of ECS is not yet elucidated in humans. ''A representation of the editing complex is shown below.'' | ||
Revision as of 21:59, 22 February 2009
Adenosine Deaminases that act on RNA (ADARs)
The Functions of the ADAR Family
- Three members of this deaminase family are known: ADAR 1, ADAR 2, and ADAR 3 that share a common modular domain structure.
- ADARs modify pre-mRNA and recognize structural determinants in the RNA.
- In mice, the editosomes with ADAR proteins require some cis-acting elements like an intronic 'editing-site complementary sequence (ECS)'. Although evolutionarily conserved, the actual role of ECS is not yet elucidated in humans. A representation of the editing complex is shown below.
- The deamination of adenosines (A) to inosine (I) is the most common editing event.
Why edit pre-mRNA?
- To increase genetic diversity through changes in the transcriptome. For an example of this, see below.
ADAR Functions in Receptors
- Deamination by editing in pre-mRNAs encodes a variety of subunits of ionotropic glutamate receptors (GluRs).
- Editing at the Q/R site of the GluR2 (GluRB) subunit of AMPA receptors changes a Gln codon CAG to an Arg codon CIG, which makes the heteromeric receptor impermeable to Ca2+ ions.
- Editing of 5-HT2C subtype serotonin receptor mRNA results in receptor isoforms with reduced G-protein coupling efficiency (reviewed by Gerber and Keller, 2001).
References
- Blanc, V, Davidson, NO. C-to-U RNA editing: mechanisms leading to genetic diversity. 2003 J Biol Chem
- Bass, BL. RNA editing by adenosine deaminases that act on RNA. 2002 Annu Rev Biochem
- Melcher, T, Maas, S, Herb, A, Sprengel, R, Higuchi, M, Seeburg, PH. RED2, a brain-specific member of the RNA-specific adenosine deaminase family. 1997 J Biol Chem
- Gerber, AP, Keller, W. RNA editing by base deamination: more enzymes, more targets, new mysteries. 2001 Trends Biochem Sci
- Polson, AG, Crain, PF, Pomerantz, SC, McCloskey, JA, Bass, BL. The mechanism of adenosine to inosine conversion by the double-stranded RNA unwinding/modifying activity: a high-performance liquid chromatography-mass spectrometry analysis. 1992 Biochemistry
- http://www.reactome.org/cgi-bin/eventbrowser