CH391L/S12/TranscriptionPromotersandTerminators: Difference between revisions
No edit summary |
No edit summary |
||
| Line 4: | Line 4: | ||
===Introduction=== | ===Introduction=== | ||
The first step toward gene expression is the transcription of a DNA template into a complementary RNA strand. This process is done by RNA polymerase, which reads the DNA template and produces an antiparallel RNA copy. As in DNA replication, the complementary strand is produced 5'->3'If the DNA template encodes for a gene, this RNA transcript will be refined into mRNA, which is further translated into a functional protein | The first step toward gene expression is the transcription of a DNA template into a complementary RNA strand. This process is done by RNA polymerase, which reads the DNA template and produces an antiparallel RNA copy. As in DNA replication, the complementary strand is produced 5'->3'If the DNA template encodes for a gene, this RNA transcript will be refined into mRNA, which is further translated into a functional protein. The RNA transcript may also go on to make ribosomal RNA (rRNA), transfer RNA (tRNA), or many other RNA products. The entire process can be broken into three major steps: initiation, elongation, and termination. | ||
==Initiation== | ==Initiation== | ||
Initiation of transcription occurs differently in eukaryotes and prokaryotes. In eukaryotes, the transcription initiation complex must be formed. This includes, the core promoter, transcription factors, RNA polymerase, and activators/repressors. In | Initiation of transcription occurs differently in eukaryotes and prokaryotes. In eukaryotes, the transcription initiation complex must be formed. This includes, the core promoter, transcription factors, RNA polymerase, and activators/repressors. In E. coli, RNA polymerase and sigma factors are needed, as well, it may be necessary to have activators/repressors based on the promoter being used. For E. coli, the RNA polymerase will bind tightly to the promoter to form an open promoter complex, then must choose the transcription start site and escape from the promoter. It is necessary to balance the strength of promoter binding the ability to escape so elongation can happen. RNAP may undergo abortive initiation in which it will form many short 9-10 bp segments until it clears the promoter and begins elongation. | ||
=='''Promoters'''== | =='''Promoters'''== | ||
A DNA sequence that recruits transcriptional machinery and lead to transcription of downstream DNA. In | A DNA sequence that recruits transcriptional machinery and lead to transcription of downstream DNA. In E. coli the -10bp and -35bp are locations of the most well conserved DNA sequences in bacterial promoters. There are on average 17 bp between the two sequences and 7 bp between the -10bp location and the transcription start site. Consensus sequences are the nucleotide sequences that share a common function, which is binding to RNAP in the case of promoters. Promoters that most closely resemble the consensus sequence will be the strongest promoters, just as those that differ from the consensus sequence will be weaker promoters. Interestingly, there has not been a promoter found in E. coli that is of the consesus sequence, it would likely bind so strongly that elongation would not occur. | ||
[[Image:PromoterIcon.png|thumb|400px|right]] | [[Image:PromoterIcon.png|The consensus Sequence of E. coli shown in lavender box|thumb|400px|right]] | ||
====Constitutive==== | ====Constitutive==== | ||
| Line 29: | Line 29: | ||
===#$% Yeast Promoter=== | ===#$% Yeast Promoter=== | ||
=== | ===Strong Bacterial Promoter LacUV5=== | ||
==Termination== | ==Termination== | ||
[[Image:500px-RegistryTerminator.jpg|thumb|400px|right]] | [[Image:500px-RegistryTerminator.jpg|thumb|400px|right]] | ||
| Line 48: | Line 48: | ||
===#$% Yeast Terminator=== | ===#$% Yeast Terminator=== | ||
==References== | |||
<biblio> | |||
#Carpousis1984 pmid=2409292 | |||
#NoelRJ2000 pmid=10713082 | |||
Revision as of 11:31, 26 February 2012
Transcription
Introduction
The first step toward gene expression is the transcription of a DNA template into a complementary RNA strand. This process is done by RNA polymerase, which reads the DNA template and produces an antiparallel RNA copy. As in DNA replication, the complementary strand is produced 5'->3'If the DNA template encodes for a gene, this RNA transcript will be refined into mRNA, which is further translated into a functional protein. The RNA transcript may also go on to make ribosomal RNA (rRNA), transfer RNA (tRNA), or many other RNA products. The entire process can be broken into three major steps: initiation, elongation, and termination.
Initiation
Initiation of transcription occurs differently in eukaryotes and prokaryotes. In eukaryotes, the transcription initiation complex must be formed. This includes, the core promoter, transcription factors, RNA polymerase, and activators/repressors. In E. coli, RNA polymerase and sigma factors are needed, as well, it may be necessary to have activators/repressors based on the promoter being used. For E. coli, the RNA polymerase will bind tightly to the promoter to form an open promoter complex, then must choose the transcription start site and escape from the promoter. It is necessary to balance the strength of promoter binding the ability to escape so elongation can happen. RNAP may undergo abortive initiation in which it will form many short 9-10 bp segments until it clears the promoter and begins elongation.
Promoters
A DNA sequence that recruits transcriptional machinery and lead to transcription of downstream DNA. In E. coli the -10bp and -35bp are locations of the most well conserved DNA sequences in bacterial promoters. There are on average 17 bp between the two sequences and 7 bp between the -10bp location and the transcription start site. Consensus sequences are the nucleotide sequences that share a common function, which is binding to RNAP in the case of promoters. Promoters that most closely resemble the consensus sequence will be the strongest promoters, just as those that differ from the consensus sequence will be weaker promoters. Interestingly, there has not been a promoter found in E. coli that is of the consesus sequence, it would likely bind so strongly that elongation would not occur.
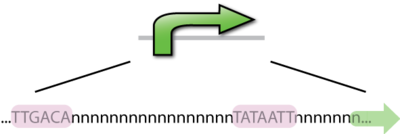
Constitutive
An unregulated promoter that allows for continual transcription of its associated genes. These promoters do not rely on input and depend only the level of free RNA polymerase holoenzyme are referred to as constitutive. Since the holoenzyme is needed, it can also be said that these rely on the level of sigma factors.
Positively
These promoters depend on the level of transcription factors that are not sigma factors. As the concentration of activator increase, the rate of transcription also increases. If an activator protein relies on the binding of an exogenous molecule to activate it, then the promoter may be referred to as inducible.
Negatively
Negatively regulated promoters on the level of a repressor transcription factor. Increased levels of a repressor will lower the activity of these promoters. If a repressor that inactivates the promoter is always present and an exogenous molecule is added that binds the repressor and deactivates it, then promoter may be referred to as inducible.
Multi-regulated
Promoters in this category are either positively or negatively regulated by multiple transcription factors. These are most useful when a promoter that relies on multiple environmental factors to function is desired.
Sigma Factors
#$% Yeast Promoter
Strong Bacterial Promoter LacUV5
Termination

The termination step of transcription varies between prokaryotes and eukaryotes. In prokaryotic organisms, termination involves formation of a hairpin loop which destabilizes the transcript in rho-independent termination. In rho-dependent termination, a protein Rho destabilizes the RNA-DNA complex and leads to termination of transcription.
Prokaryotic
Rho-Dependent
In this type of termination, a protein factor called Rho destabilizes the DNA template-RNA transcript complex, causing the release of the RNA transcript. Rho-dependent terminators are not included in the iGEM registry because these terminators are not specified by sequence.
Rho-Independent
The terminators are composed sequences that lead to a hairpin loop rich in G-C base pairs followed by many bases of thymine. The formation of the RNA G-C rich stem loop causes a pause in the RNA Polymerase. This pause, followed by the transcription of the poly A tail into a run of U's causes a mechanical stress and the unwinding of the RNA-DNA complex, causing the dissociation of the RNA transcript from RNA polymerase.
#$% Bacterial Terminator
Eukaryotic
In yeast, termination is different for each RNA polymerase (I-III). The process involves the polyadenylation at the 3' end of the RNA transcript. A set of proteins cleave off the RNA transcript and then synthesize the poly A tail, independent of the DNA template. This step is important toward refining the RNA into mRNA that will translated.
#$% Yeast Terminator
References
<biblio>
- Carpousis1984 pmid=2409292
- NoelRJ2000 pmid=10713082