CH391L/S13/DirectedProteinEvolution: Difference between revisions
No edit summary |
|||
| (15 intermediate revisions by 2 users not shown) | |||
| Line 2: | Line 2: | ||
'''[http://en.wikipedia.org/wiki/Directed_evolution Directed evolution]''' is a powerful method for altering the properties of biological parts and systems. '''Directed protein evolution''' employs iterative rounds of mutation and artificial selection to generate new proteins with desirable functions <cite>Romero2009</cite>. | '''[http://en.wikipedia.org/wiki/Directed_evolution Directed evolution]''' is a powerful method for altering the properties of biological parts and systems. '''Directed protein evolution''' employs iterative rounds of mutation and artificial selection to generate new proteins with desirable functions <cite>Romero2009</cite>. | ||
== Introduction == | == Introduction == | ||
[[Image:Overview_of_directed_evolution.jpg | thumb | right | 200 px | Overview of directed evolution <cite>Romero2009</cite>.]] | |||
Biological molecules have a unique ability to rapidly evolve in response to strong selective pressure <cite>Romero2009</cite>. Protein engineers exploit this evolvability to generate new and useful proteins through successive rounds of mutation and selection <cite>Stemmer1995</cite> <cite>Giver1998</cite>. This approach is known as directed protein evolution, and it involves four basic steps <cite>Romero2009</cite>: | Biological molecules have a unique ability to rapidly evolve in response to strong selective pressure <cite>Romero2009</cite>. Protein engineers exploit this evolvability to generate new and useful proteins through successive rounds of mutation and selection <cite>Stemmer1995</cite> <cite>Giver1998</cite>. This approach is known as directed protein evolution, and it involves four basic steps <cite>Romero2009</cite>: | ||
#A parent protein sequence is selected. | |||
#The parent sequence is mutated to generate a library of functional variants. | |||
#Variants are evaluated for their ability to perform the desired function. | |||
#The process is repeated until the desired function is achieved. | |||
The parent sequence is chosen based on its perceived similarity to the desired function, and a library of functional variants is generated using one or several of a variety of sequence diversification techniques <cite>Romero2009</cite>. High-throughput functional screens and genetic selection methods are used to identify library members with enhanced target function, and those variants are used as parent sequences in successive rounds of mutation and selection <cite>Romero2009</cite>. This process is repeated until the desired function is achieved. | The parent sequence is chosen based on its perceived similarity to the desired function, and a library of functional variants is generated using one or several of a variety of sequence diversification techniques <cite>Romero2009</cite>. High-throughput functional screens and genetic selection methods are used to identify library members with enhanced target function, and those variants are used as parent sequences in successive rounds of mutation and selection <cite>Romero2009</cite>. This process is repeated until the desired function is achieved. | ||
== Library construction == | == Library construction == | ||
| Line 20: | Line 24: | ||
'''Error-prone PCR''' is a technique for introducing random point mutations into cloned sequences, in which modifications to standard PCR conditions increase the error rate of nucleotide incorporation during amplification <cite>Cadwell1992</cite> <cite>Abou-Nader2010</cite>. Common methods for decreasing polymerase fidelity include the addition of manganese ions, an increase in the concentration of magnesium, and using an imbalanced ration of dNTPs <cite>Cadwell1992</cite> <cite>Abou-Nader2010</cite>. There are a number of commercially available kits for error-prone PCR, such as the GeneMorph II random mutagenesis kit from [http://www.genomics.agilent.com/CollectionSubpage.aspx?PageType=Product&SubPageType=ProductData&PageID=376 Agilent Technologies]. | '''Error-prone PCR''' is a technique for introducing random point mutations into cloned sequences, in which modifications to standard PCR conditions increase the error rate of nucleotide incorporation during amplification <cite>Cadwell1992</cite> <cite>Abou-Nader2010</cite>. Common methods for decreasing polymerase fidelity include the addition of manganese ions, an increase in the concentration of magnesium, and using an imbalanced ration of dNTPs <cite>Cadwell1992</cite> <cite>Abou-Nader2010</cite>. There are a number of commercially available kits for error-prone PCR, such as the GeneMorph II random mutagenesis kit from [http://www.genomics.agilent.com/CollectionSubpage.aspx?PageType=Product&SubPageType=ProductData&PageID=376 Agilent Technologies]. | ||
'''[http://en.wikipedia.org/wiki/DNA_shuffling DNA shuffling]''' is a technique for | '''[http://en.wikipedia.org/wiki/DNA_shuffling DNA shuffling]''' is a technique for “''in vitro'' homologous recombination of pools of selected mutant genes" <cite>Stemmer1994</cite>. In this method, parental sequences are fragmented by DNase I and then reassembled by PCR. Recombination events occur as fragments anneal at regions of sufficient sequence homology <cite>Stemmer1994</cite>. After reassembly, chimeric sequences are amplified by PCR and cloned into an appropriate vector. | ||
== Screening and selection == | == Screening and selection == | ||
Once a library of mutants is generated, they must be evaluated for their ability to perform the desired function. To do this, protein engineers employ a variety of high-throughput assays. Successful assays allow | Once a library of mutants is generated, they must be evaluated for their ability to perform the desired function. To do this, protein engineers employ a variety of high-throughput assays. Successful assays allow researchers to test a large number of functional variants while maintaining a connection between phenotype (the evolved protein function) and genotype (the DNA sequence encoding the evolved function) <cite>Lin2002</cite>. These assays can be categorized as either “''in vivo''” versus “''in vitro''” or “selection” versus “screening" <cite>Leemhuis2009</cite>. The most important distinction is between a screen and a selection. Selections allow for only cells expressing proteins that exhibit the desired function to survive. In contrast, a screen allows for cells expressing any functional variant to survive yet be distinguished by phenotype. In a typical screen, the number of variants that can be tested is ~10<sup>4</sup> <cite>Leemhuis2009</cite>. This is often due to the fact that researchers must pick individual colonies to grow liquid cultures and personally supervise activity assays. In a typical selection, the number of variants that can be tested is on the order of ~10<sup>6</sup> to 10<sup>8</sup> <cite>Leemhuis2009</cite>. Selections are limited by transformation efficiency, which is ~10<sup>6</sup> in yeast and ~10<sup>8</sup> in ''E. coli''. | ||
The most basic cell-based screening methods involve transforming a library of mutants into bacteria and identifying individual colonies or cultures that exhibit the desired function. These assays maintain a link between phenotype and genotype that is “achieved naturally by introducing plasmid DNA encoding the protein into a cell | The most basic cell-based screening methods involve transforming a library of mutants into bacteria and identifying individual colonies or cultures that exhibit the desired function. These assays maintain a link between phenotype and genotype that is “achieved naturally by introducing plasmid DNA encoding the protein into a cell" <cite>Lin2002</cite> These methods allow millions of sequence variants to be transformed into cells, and by manipulating the statistic of DNA transformation allow for each cell to contain a single vector containing a single sequence variant. These individual cells can then be isolated by growth on solid media <cite>Lin2002</cite>. | ||
'''List of more advanced high-throughput screening and selection methods:''' | |||
#[http://en.wikipedia.org/wiki/Phage_display phage display] | |||
#[http://en.wikipedia.org/wiki/Ribosome_display ribosome display] | |||
#[http://www.pnas.org/content/94/23/12297.short mRNA-peptide fusion] | |||
#[http://pubs.acs.org/doi/abs/10.1021/cb100423u plasmid display] | |||
#[http://www.sciencedirect.com/science/article/pii/S0167779902000069 cell-surface display] | |||
#[http://www.nature.com/nature/journal/v380/n6574/abs/380548a0.html n-hybrid systems] | |||
#[http://www.nature.com/nmeth/journal/v3/n7/full/nmeth897.html ''in vitro'' compartmentalization] | |||
#[http://www.nature.com/nbt/journal/v18/n4/full/nbt0400_393.html spatial address] | |||
== Improving whole cell fluorescence of GFP by directed evolution == | == Improving whole cell fluorescence of GFP by directed evolution == | ||
Wild type green fluorescent protein (GFP) is routinely used as a reporter of gene regulation. However, for some applications, a stronger whole cell fluorescence signal is required. Thus, the research team of Willem Stemmer et al. set out to construct a GFP mutant that would exhibit enhanced whole cell fluorescence. To do this, the group first constructed a synthetic GFP gene with improved codon usage. They then performed successive cycles of DNA shuffling followed by a visual screen for the brightest E. coli colonies. Using this method, they were able to generate a mutant with a whole cell fluorescence signal 45-fold greater than a standard commercially available GFP sequence. | [[Image:Comparison_of_GFP_flurorescence.jpg | thumb | right | 200 px | Comparison of the fluorescence of different GFP constructs in whole ''E. coli'' cells <cite>Stemmer1995</cite>.]] | ||
Wild type green fluorescent protein (GFP) is routinely used as a reporter of gene regulation. However, for some applications, a stronger whole cell fluorescence signal is required <cite>Stemmer1995</cite>. Thus, the research team of Willem Stemmer et al. set out to construct a GFP mutant that would exhibit enhanced whole cell fluorescence. To do this, the group first constructed a synthetic GFP gene with improved [http://en.wikipedia.org/wiki/Codon_usage_bias codon usage]. They then performed successive cycles of DNA shuffling followed by a visual screen for the brightest ''E. coli'' colonies. Using this method, they were able to generate a mutant with a whole cell fluorescence signal 45-fold greater than a standard commercially available GFP sequence <cite>Stemmer1995</cite>. | |||
== Directed evolution of a thermostable esterase == | == Directed evolution of a thermostable esterase == | ||
It had been previously suggested that enzyme thermostability is incompatible with high catalytic activity at low temperature. However, by constraining both properties, the Frances Arnold group was able to create a thermostable esterase that maintains high catalytic activity at lower temperatures using directed protein evolution. To do this, the Arnold group generated a mutant library through error-prone PCR and DNA shuffling and developed a 96-well plate thermostability and activity screen. After six generations of mutagenesis and screening they were able to generate a thermostable p-nitrobenzyl esterase mutant that retains catalytic activity at low temperature. | [[Image:model_of_pNB_esterase.jpg | thumb | left | 200 px | Model of pNB esterase constructed based on homology to esterases of known structures <cite>Giver1998</cite>.]] | ||
It had been previously suggested that enzyme thermostability is incompatible with high catalytic activity at low temperature <cite>Giver1998</cite>. However, by constraining both properties, the Frances Arnold group was able to create a thermostable esterase that maintains high catalytic activity at lower temperatures using directed protein evolution. To do this, the Arnold group generated a mutant library through error-prone PCR and DNA shuffling and developed a 96-well plate thermostability and activity screen. After six generations of mutagenesis and screening they were able to generate a thermostable p-nitrobenzyl esterase mutant that retains catalytic activity at low temperature <cite>Giver1998</cite>. | |||
== iGEM connection == | == iGEM connection == | ||
The UC Davis [http://2012.igem.org/Team:UC_Davis/Project 2012 iGEM team] used directed evolution to engineer an E. coli strain that more efficiently degrades ethylene glycol than previous strains. Although not strictly a directed protein evolution project, their work demonstrates the ability of biological molecules and systems to rapidly evolve under strong selective pressure. | The UC Davis [http://2012.igem.org/Team:UC_Davis/Project 2012 iGEM team] used directed evolution to engineer an E. coli strain that more efficiently degrades ethylene glycol than previous strains. Although not strictly a directed protein evolution project, their work demonstrates the ability of biological molecules and systems to rapidly evolve under strong selective pressure. | ||
==References== | ==References== | ||
<biblio> | <biblio> | ||
#Romero2009 Romero | #Romero2009 Romero PA and Arnold FH. Exploring protein fitness landscapes by directed evolution. Nat Rev Mol Cell Bio, 2009. | ||
#Stemmer1995 Crameri A, Whitehorn EA, Tate E, Stemmer WP. Improved green fluorescent protein by molecular evolution using DNA shuffling. Nat Biotechnol, 1996. | #Stemmer1995 Crameri A, Whitehorn EA, Tate E, Stemmer WP. Improved green fluorescent protein by molecular evolution using DNA shuffling. Nat Biotechnol, 1996. | ||
#Giver1998 Giver L, Gershenson A, Freskgard PO, Arnold FH. Directed evolution of a thermostable esterase. Proc Natl Acad Sci USA, 1998. | #Giver1998 Giver L, Gershenson A, Freskgard PO, Arnold FH. Directed evolution of a thermostable esterase. Proc Natl Acad Sci USA, 1998. | ||
#Cadwell1992 Cadwell RC and Joyce GF. Randomization of genes by PCR mutagenesis. Genome Res, 1992. | #Cadwell1992 Cadwell RC and Joyce GF. Randomization of genes by PCR mutagenesis. Genome Res, 1992. | ||
#Abou-Nader2010 Abou-Nader M and Benedik MJ. Rapid generation of random mutant libraries. Bioeng Bugs, 2010. | #Abou-Nader2010 Abou-Nader M and Benedik MJ. Rapid generation of random mutant libraries. Bioeng Bugs, 2010. | ||
#Stemmer1994 Stemmer WP. Rapid evolution of a protein by in vitro DNA shuffling. Nature, 1994. | #Stemmer1994 Stemmer WP. Rapid evolution of a protein by ''in vitro'' DNA shuffling. Nature, 1994. | ||
#Lin2002 Lin H and Cornish VW. Screening and selection methods for large-scale analysis of protein function. | #Lin2002 Lin H and Cornish VW. Screening and selection methods for large-scale analysis of protein function. | ||
#Leemhuis2009 Leemhuis H, Kelly RM, Dijkhuizen L. Directed evolution of enzymes: library screening strategies. IUBMB Life, 2009. | |||
</biblio> | </biblio> | ||
Revision as of 12:39, 4 March 2013
What is directed protein evolution?
Directed evolution is a powerful method for altering the properties of biological parts and systems. Directed protein evolution employs iterative rounds of mutation and artificial selection to generate new proteins with desirable functions [1].
Introduction

Biological molecules have a unique ability to rapidly evolve in response to strong selective pressure [1]. Protein engineers exploit this evolvability to generate new and useful proteins through successive rounds of mutation and selection [2] [3]. This approach is known as directed protein evolution, and it involves four basic steps [1]:
- A parent protein sequence is selected.
- The parent sequence is mutated to generate a library of functional variants.
- Variants are evaluated for their ability to perform the desired function.
- The process is repeated until the desired function is achieved.
The parent sequence is chosen based on its perceived similarity to the desired function, and a library of functional variants is generated using one or several of a variety of sequence diversification techniques [1]. High-throughput functional screens and genetic selection methods are used to identify library members with enhanced target function, and those variants are used as parent sequences in successive rounds of mutation and selection [1]. This process is repeated until the desired function is achieved.
Library construction
After a parent sequence is chosen, a library of functional mutants must be generated. Common methods used for library construction include error-prone PCR and DNA shuffling [4] [5].
Error-prone PCR is a technique for introducing random point mutations into cloned sequences, in which modifications to standard PCR conditions increase the error rate of nucleotide incorporation during amplification [4] [5]. Common methods for decreasing polymerase fidelity include the addition of manganese ions, an increase in the concentration of magnesium, and using an imbalanced ration of dNTPs [4] [5]. There are a number of commercially available kits for error-prone PCR, such as the GeneMorph II random mutagenesis kit from Agilent Technologies.
DNA shuffling is a technique for “in vitro homologous recombination of pools of selected mutant genes" [6]. In this method, parental sequences are fragmented by DNase I and then reassembled by PCR. Recombination events occur as fragments anneal at regions of sufficient sequence homology [6]. After reassembly, chimeric sequences are amplified by PCR and cloned into an appropriate vector.
Screening and selection
Once a library of mutants is generated, they must be evaluated for their ability to perform the desired function. To do this, protein engineers employ a variety of high-throughput assays. Successful assays allow researchers to test a large number of functional variants while maintaining a connection between phenotype (the evolved protein function) and genotype (the DNA sequence encoding the evolved function) [7]. These assays can be categorized as either “in vivo” versus “in vitro” or “selection” versus “screening" [8]. The most important distinction is between a screen and a selection. Selections allow for only cells expressing proteins that exhibit the desired function to survive. In contrast, a screen allows for cells expressing any functional variant to survive yet be distinguished by phenotype. In a typical screen, the number of variants that can be tested is ~104 [8]. This is often due to the fact that researchers must pick individual colonies to grow liquid cultures and personally supervise activity assays. In a typical selection, the number of variants that can be tested is on the order of ~106 to 108 [8]. Selections are limited by transformation efficiency, which is ~106 in yeast and ~108 in E. coli.
The most basic cell-based screening methods involve transforming a library of mutants into bacteria and identifying individual colonies or cultures that exhibit the desired function. These assays maintain a link between phenotype and genotype that is “achieved naturally by introducing plasmid DNA encoding the protein into a cell" [7] These methods allow millions of sequence variants to be transformed into cells, and by manipulating the statistic of DNA transformation allow for each cell to contain a single vector containing a single sequence variant. These individual cells can then be isolated by growth on solid media [7].
List of more advanced high-throughput screening and selection methods:
- phage display
- ribosome display
- mRNA-peptide fusion
- plasmid display
- cell-surface display
- n-hybrid systems
- in vitro compartmentalization
- spatial address
Improving whole cell fluorescence of GFP by directed evolution

Wild type green fluorescent protein (GFP) is routinely used as a reporter of gene regulation. However, for some applications, a stronger whole cell fluorescence signal is required [2]. Thus, the research team of Willem Stemmer et al. set out to construct a GFP mutant that would exhibit enhanced whole cell fluorescence. To do this, the group first constructed a synthetic GFP gene with improved codon usage. They then performed successive cycles of DNA shuffling followed by a visual screen for the brightest E. coli colonies. Using this method, they were able to generate a mutant with a whole cell fluorescence signal 45-fold greater than a standard commercially available GFP sequence [2].
Directed evolution of a thermostable esterase
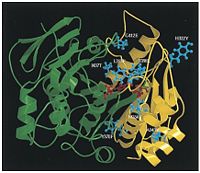
It had been previously suggested that enzyme thermostability is incompatible with high catalytic activity at low temperature [3]. However, by constraining both properties, the Frances Arnold group was able to create a thermostable esterase that maintains high catalytic activity at lower temperatures using directed protein evolution. To do this, the Arnold group generated a mutant library through error-prone PCR and DNA shuffling and developed a 96-well plate thermostability and activity screen. After six generations of mutagenesis and screening they were able to generate a thermostable p-nitrobenzyl esterase mutant that retains catalytic activity at low temperature [3].
iGEM connection
The UC Davis 2012 iGEM team used directed evolution to engineer an E. coli strain that more efficiently degrades ethylene glycol than previous strains. Although not strictly a directed protein evolution project, their work demonstrates the ability of biological molecules and systems to rapidly evolve under strong selective pressure.
References
-
Romero PA and Arnold FH. Exploring protein fitness landscapes by directed evolution. Nat Rev Mol Cell Bio, 2009.
-
Crameri A, Whitehorn EA, Tate E, Stemmer WP. Improved green fluorescent protein by molecular evolution using DNA shuffling. Nat Biotechnol, 1996.
-
Giver L, Gershenson A, Freskgard PO, Arnold FH. Directed evolution of a thermostable esterase. Proc Natl Acad Sci USA, 1998.
-
Cadwell RC and Joyce GF. Randomization of genes by PCR mutagenesis. Genome Res, 1992.
-
Abou-Nader M and Benedik MJ. Rapid generation of random mutant libraries. Bioeng Bugs, 2010.
-
Stemmer WP. Rapid evolution of a protein by in vitro DNA shuffling. Nature, 1994.
-
Lin H and Cornish VW. Screening and selection methods for large-scale analysis of protein function.
-
Leemhuis H, Kelly RM, Dijkhuizen L. Directed evolution of enzymes: library screening strategies. IUBMB Life, 2009.