Frankel:Force Spectroscopy
<owwmenu align="center" font="helvetica" bold="1" color="white" bgcolor="black" hovercolor="black" bghovercolor="orange" topfontsize="10" fontSize="10" image="Danbanner-bio-machines.jpg" >
Home=Frankel
Members=#,Principal Investigator=Frankel:Lab_Members, PhD students=Frankel:Lab_Members, Alumni=Frankel:Lab_Members
Contact=Frankel:Contact
Collaborators=Frankel:Collaborators
Publications=Frankel:Publications
Lab=Frankel:Research
Research=#,Force Spectroscopy=Frankel:Force Spectroscopy,HIV/Virus=Frankel:HIV/Virus,ECM Proteins=Frankel:ECM Proteins,Cyberplasm=Frankel:Cyberplasm,Cancer=Frankel:Cancer
Force Spectroscopy
|
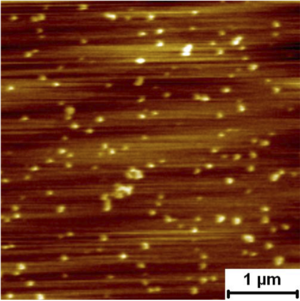
Assembly Self assembly and pore formation of HIV GP160 revealed at molecular resolution GP160mica Force spectra taken on raised terraces and lower features. Rupture force distribution of self assembled gp160 unfolding on terraced and lower regions. Rupture forces were measured as 79.6 ± 3.9 pN and 81.3 ± 3.8 pN for the terraces and lower regions, respectively. These forces are much lower than those measured for unfolding of isolated proteins on mica, which were above 160 pN. The lower unfolding forces suggest that GP160 is considerably easier to unfold when aggregated than isolated.
| ||
|
Revealing the selective interactions of fibronectin with lipid bilayer.
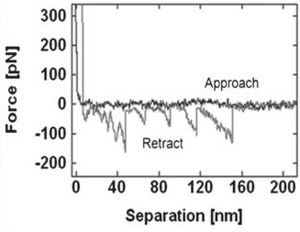
Sawtooth pattern on the retraction force curve indicating the unfolding of fibronectin. The average rupture force distribution of the protein on mica surface was 85.1 ± 2.7 pN. |