Haynes:NewProtocol: Difference between revisions
(New page: New Protocol) |
Rene M Davis (talk | contribs) |
||
| (5 intermediate revisions by the same user not shown) | |||
| Line 1: | Line 1: | ||
<- [[Haynes:Protocols | Back to Protocols]] | |||
<div style="width: 800px" align="center"> | |||
<font size=3>'''Gibson Assembly'''</font><br> | |||
By René Davis, 2012</div> | |||
<br> | |||
==Overview== | |||
<div style="width: 800px"> | |||
The Gibson Assembly method allows you to assemble multiple DNA parts in just 2 steps.<br> | |||
In the first step, 20 basepair overlaps are added to the sequences to be assembled: | |||
[[image:Slide1.png|500px|Gibson Assembly step 1]] <br> | |||
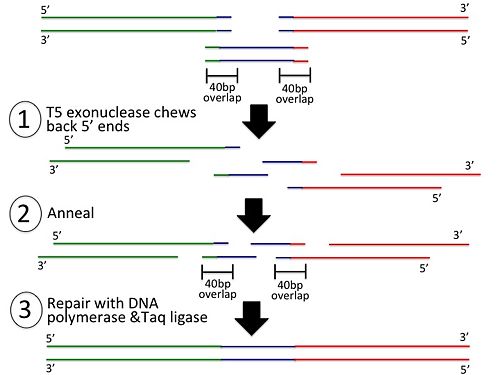
In the second step, the PCR products are added to the Gibson Assembly Mix: | |||
[[image:Slide2.jpg|500px|Gibson Assembly step 2]] | |||
==Materials== | |||
Designing primers: | |||
[[Image:Slide_3_pdf.png|500px|Gibson Assembly primers]]<br> | |||
PCR mix<br> | |||
15ul of Gibson Assembly Mix | |||
(I will add the recipe soon, but for now, find 15ul aliquotes in PCR tubes in a clearly labeled bag in the -20C freezer door... soonish) | |||
==Procedure== | |||
# In a PCR tube, mix the components on ice in the order they are listed above. | |||
# Perform thermocycling program | |||
## | |||
For now: http://django.gibthon.org/help/gibson/ | |||
==Notes== | |||
Please feel free to post comments, questions, or improvements to this protocol. Happy to have your input! | |||
#List troubleshooting tips here. | |||
#You can also link to FAQs/tips provided by other sources such as the manufacturer or other websites. | |||
#Anecdotal observations that might be of use to others can also be posted here. | |||
Please sign your name to your note by adding <font face="courier"><nowiki>'''*~~~~''':</nowiki></font> to the beginning of your tip. | |||
==References== | |||
'''Relevant papers and books''' | |||
<!-- If this protocol has papers or books associated with it, list those references here.--> | |||
<!-- See the [[OpenWetWare:Biblio]] page for more information, but this format doesn't work currently. | |||
<biblio> | |||
#Goldbeter-PNAS-1981 pmid=6947258 | |||
#Jacob-JMB-1961 pmid=13718526 | |||
#Ptashne-Genetic-Switch isbn=0879697164 | |||
</biblio>--> | |||
<!-- Try the [[Template:FormatRef|FormatRef template]]--> | |||
#{{FormatRef|Gibson, DG, Young L, Chuang RY, Venter JC, Hutchison CA, Smith HO |2009| |Nature Methods 6(5)|343-5| }} PMID 19363495 | |||
==Contact== | |||
*René at rene.davis at asu dot edu | |||
or instead, [[Talk:{{PAGENAME}}|discuss this protocol]]. | |||
<!-- You can tag this protocol with various categories. See the [[Categories]] page for more information. --> | |||
<!-- Move the relevant categories above this line to tag your protocol with the label | |||
[[Category:Protocol]] | |||
[[Category:Needs attention]] | |||
[[Category:In vitro]] | |||
[[Category:In vivo]] | |||
[[Category:In silico]] | |||
[[Category:DNA]] | |||
[[Category:RNA]] | |||
[[Category:Protein]] | |||
[[Category:Chemical]] | |||
[[Category:Escherichia coli]] | |||
[[Category:Yeast]] | |||
--> | |||
Latest revision as of 13:04, 26 October 2012
Gibson Assembly
Overview
The Gibson Assembly method allows you to assemble multiple DNA parts in just 2 steps.
In the first step, 20 basepair overlaps are added to the sequences to be assembled:
In the second step, the PCR products are added to the Gibson Assembly Mix:
Materials
Designing primers:
PCR mix
15ul of Gibson Assembly Mix
(I will add the recipe soon, but for now, find 15ul aliquotes in PCR tubes in a clearly labeled bag in the -20C freezer door... soonish)
Procedure
- In a PCR tube, mix the components on ice in the order they are listed above.
- Perform thermocycling program
For now: http://django.gibthon.org/help/gibson/
Notes
Please feel free to post comments, questions, or improvements to this protocol. Happy to have your input!
- List troubleshooting tips here.
- You can also link to FAQs/tips provided by other sources such as the manufacturer or other websites.
- Anecdotal observations that might be of use to others can also be posted here.
Please sign your name to your note by adding '''*~~~~''': to the beginning of your tip.
References
Relevant papers and books
- Gibson, DG, Young L, Chuang RY, Venter JC, Hutchison CA, Smith HO (2009) - Nature Methods 6(5) 343-5 PMID 19363495
Contact
- René at rene.davis at asu dot edu
or instead, discuss this protocol.