IGEM:Caltech/2007/Project/N: Difference between revisions
From OpenWetWare
Jump to navigationJump to search
| (One intermediate revision by the same user not shown) | |||
| Line 15: | Line 15: | ||
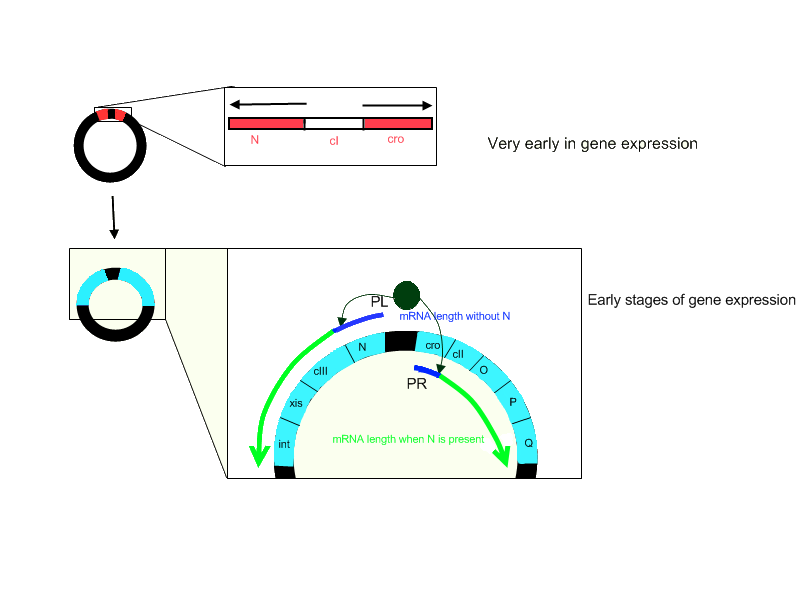
N protein is one of the three critical proteins expressed by the lambda genome that influence the developmental cycles of the virus. It is called an antiterminator because in the presence of this protein, RNA polymerase is able to code through regions of the genome it would otherwise be unable to transcribe. As depicted in the diagram below, there is a termination sequence early in the lambda genome, which causes the mRNA transcript to terminate in the absence of N. However, when N is present, the protein recognizes a specific sequence called Nut, which stands for N utilization, and as the polymerase passes over the Nut site, it is modified by N so that it can ignore the termination sequence that is downstream of the Nut site, thus allowing the elongation of the mRNA transcript. | N protein is one of the three critical proteins expressed by the lambda genome that influence the developmental cycles of the virus. It is called an antiterminator because in the presence of this protein, RNA polymerase is able to code through regions of the genome it would otherwise be unable to transcribe. As depicted in the diagram below, there is a termination sequence early in the lambda genome, which causes the mRNA transcript to terminate in the absence of N. However, when N is present, the protein recognizes a specific sequence called Nut, which stands for N utilization, and as the polymerase passes over the Nut site, it is modified by N so that it can ignore the termination sequence that is downstream of the Nut site, thus allowing the elongation of the mRNA transcript. | ||
[[Image:N_antiterminator.png | [[Image:N_antiterminator.png]] | ||
==Role in Lambda Life Cycle== | ==Role in Lambda Life Cycle== | ||
| Line 24: | Line 24: | ||
The objective of this subproject project was to determine the necessary concentration of N protein required for lysis. The N protein is active in the early stages of viral development, even before a choice is made between lysis and lysogeny, so in the absence of this protein the virus will not be infectious. | The objective of this subproject project was to determine the necessary concentration of N protein required for lysis. The N protein is active in the early stages of viral development, even before a choice is made between lysis and lysogeny, so in the absence of this protein the virus will not be infectious. | ||
The constructed system consists of the N gene driven by the Ptet promoter and followed by a transcription terminator (B0015). This device is followed by a sequence of either a strong or medium strength promoter (J23100 and J23116 respectively), TetR gene and terminator, which is responsible for expressing TetR protein, which is a repressor that acts on the Ptet promoter. | The constructed system, as depicted below, consists of the N gene driven by the Ptet promoter and is followed by a transcription terminator (B0015). This device is followed by a sequence of either a strong or medium strength promoter (J23100 and J23116 respectively), TetR gene and terminator, which is responsible for expressing TetR protein, which is a repressor that acts on the Ptet promoter. | ||
We designed this system so that we would be able to control the output of protein from the cell by controlling the input to the cell. Under normal conditions, this construct should stop producing N protein as the TetR protein expressed by the TetR gene represses the Ptet promoter. However, if anhydrotetracycline (aTc) is added to the system, the aTc binds to TetR, relieving the repression of the Ptet promoter and allowing the N protein to be produced. Therefore, we can control the concentration of N protein produced in the cells by adding varying amounts of aTc to the media in the plates or liquid cell cultures. | |||
[[Image:N_construct.png]] | |||
==Status and Future Plans== | ==Status and Future Plans== | ||