IGEM:Harvard/2006/Cyanobacteria: Difference between revisions
JeffreyLau (talk | contribs) |
|||
| (129 intermediate revisions by 6 users not shown) | |||
| Line 1: | Line 1: | ||
{{IGEM:/Harvard/2006/Cyanobacteria}} | |||
__TOC__ | |||
==Introduction== | |||
Welcome to the lab notebook for the Cyanobacteria oscillator project! The goal of our team, composed of four members, is to reconstruct the cyanobacterial circadian oscillator system into E. coli. Three proteins, KaiA, B, and C, have been shown to have an in-vitro phosphorylation state oscillation (Nakajima et al. 2005) by transcriptional-translational independent methods. If this system can be reconstituted in ''E. coli'', there are two important applications: | |||
#'''Synthetic Biology''': Creating a functional, oscillating set of proteins is the next logical step from the synthetic "repressilator" system engineered by Elowitz et al. (2000). Although a good proof of concept, the "repressilator" lacks the stability needed from a robust oscillator such as the naturally evolved cyanobacterial oscillator. This robust oscillator could prove useful in an eventual biocircuit. | |||
#'''Circadian Biology''': Cyanobacteria are the simplest model organisms for the study of circadian oscillation. Although circadian oscillation has been fairly well characterized, less is understood at the molecular level. By porting the oscillation system into ''E. coli'', one can begin to understand more precisely the pathways involved in the genomic oscillation of cyanobacteria. | |||
For more background information on the ciracadian system, please check out our "Literature" section. Otherwise, day-to-day work can be found under the "Lab Notebook" tab; we will post major results of our work and links to the days as they become available. If you have questions or comments, feel free to contact us: information is located at the main Harvard iGEM 2006 page. Thanks! | |||
Sincerely,<br> | |||
Zhipeng, Hetmann, Dave, and Jeff | |||
'''Update 10/27/06:''' We believe we can express the three proteins into e. coli, and that there is interaction between A+C and possible interaction between B+C. See the Lab Notebook for more information. | |||
[[Image:102706_cyanoresult.jpg]] | |||
==Outline of Findings and Signifigant Dates== | |||
*07/05/06: The incubator for growing up our cyanobacteria is complete; we have cultures growing! [[IGEM:Harvard/2006/Cyanobacteria/Notebook/2006-7-5 | Link]] | |||
*07/10/06: Some computer modeling has been done to see the effect of multiple unsyncronized clocks on phosphorylation state output. [[IGEM:Harvard/2006/Cyanobacteria/Notebook/2006-7-10 | Link]] | |||
*07/21/06: Upon having trouble with site-specific mutagenesis on the KaiA and KaiBC operons from the cyanobacterial genome, we have decided to pursue synthesis of the constructs in parallel with continued extraction attempts. [[IGEM:Harvard/2006/Cyanobacteria/Notebook/2006-7-21 | Link]] | |||
*08/01/06: Preliminary success with site-specific mutagenesis. [[IGEM:Harvard/2006/Cyanobacteria/Notebook/2006-8-1 | Link]] | |||
*08/05/06: Promoter leakness tests come out negative. May have to use low-copy plasmids if we want good control of protein expression in Top10F. [[IGEM:Harvard/2006/Cyanobacteria/Notebook/2006-8-5 | Link]] | |||
*08/11/06: We are moving to the synthetic KaiA, KaiB, and KaiC for future work. [[IGEM:Harvard/2006/Cyanobacteria/Notebook/2006-8-11 | Link]] | |||
*08/30/06: We successfully made the first construct, Lac+RBS+KaiC. [[IGEM:Harvard/2006/Cyanobacteria/Notebook/2006-8-30 | Link]] | |||
*09/01/06: Using the newly developed ligation protocol, we have successfully repeated Lac+RBS+KaiC from 08/30/06 and made Lac+RBS+KaiA. [[IGEM:Harvard/2006/Nicholas_Stroustrup%27s_Notebook#Results_Summary | | |||
Link]] | |||
*10/21/06: Successfully made Lac+RBS+KaiB and Lac+RBS+KaiA+Lac+RBS+KaiC. [[IGEM:Harvard/2006/Cyanobacteria/Notebook/2006-10-21 | Link]] | |||
*10/24/06: Successfully made Lac+RBS+KaiB+Lac+RBS+KaiC. [[IGEM:Harvard/2006/Cyanobacteria/Notebook/2006-10-24 | Link]] | |||
*10/25/06: Constructs for Stage I have been completed; ready to move to Stage I of Western Blotting, to verify expression of KaiC and interaction of KaiA and KaiB with KaiC. [[IGEM:Harvard/2006/Cyanobacteria/Notebook/2006-10-24 | Link]] | |||
*'''10/27/06: Preliminary data indicates that the Kai proteins are being expressed in e. coli and that there is interaction between the three proteins! [[IGEM:Harvard/2006/Cyanobacteria/Notebook/2006-10-27 | Link]]''' | |||
== | ==Construct Planning== | ||
[[Image:construct_plans.png|thumb|left|330px|Constructs we plan to create.]] | |||
<br style="clear:both"> | |||
< | |||
== | === Lengths === | ||
From VF2 to VR (BioBrick primers): | |||
* KaiA + J04500: 1406 bp | |||
* KaiB + J04500: 859 bp | |||
* KaiC + J04500: 2110 bp | |||
==Agenda== | |||
''See image at right for our long-term project outline.'' | |||
[[Image:Cyanobacteria_Flowchart.png|thumb|Long-term project outline]] | |||
== | |||
'' | |||
[[Image: | |||
==BioBricks Used== | |||
:*<bbpart>BBa_J04450</bbpart> | |||
:**RFP device | |||
:**Insert size: 1069bp | |||
:**[[http://parts.mit.edu/registry/index.php/Part:pSB1A2 pSB1A2]] | |||
:***High-copy, Amp<sup>R</sup> | |||
:***Size: 2079bp | |||
:*<bbpart>BBa_J04500</bbpart> | |||
:**Lac promoter + RBS | |||
:**Insert size: 220bp | |||
:**[[http://parts.mit.edu/registry/index.php/Part:pSB1AK3 pSB1AK3]] | |||
:***High-copy, Amp<sup>R</sup>, Kan<sup>R</sup> | |||
:***Insert size: 3189bp | |||
:*[[http://parts.mit.edu/registry/index.php/Part:pSB4A3 pSB4A3]] | |||
:**Low-copy, Amp<sup>R</sup> | |||
:**Insert size: 3339 bp | |||
:*<bbpart>BBa_R0010</bbpart> + <bbpart>BBa_E0241</bbpart> | |||
:**GFP device | |||
:**Insert size: 995 bp | |||
==Presentations== | ==Presentations== | ||
*[[IGEM:Harvard/2006/Presentation_cyano_week2 | Project proposal (week 2)]] | *[[IGEM:Harvard/2006/Presentation_cyano_week2 | Project proposal (week 2)]] | ||
*[[ | *[[Media:Cyan_week3.ppt |Week 3 progress update]] | ||
**Built incubator and obtained WH8102, PCC7942, and PCC6803 strains | **Built incubator and obtained WH8102, PCC7942, and PCC6803 strains | ||
*[[Media:Cyano_week4.ppt |Week 4 progress update]] | |||
*[[Media:Cyanobacteria_Presentation_Week_5.ppt |Week 5 progress update, upd. 10:10 7/17]] | |||
*[[Media:Cyanobacteria_Presentation_Week_6.ppt |Week 6 progress update, upd. 10:02 7/24 HH]] | |||
*[[Media:Cyanobacteria_Presentation_Week_7.ppt |Week 7 progress update]] | |||
*[[Media:Cyanobacteria_presentation_Week_8.ppt |Week 8 progress update]] | |||
*[[Media:Cyanobacteria_presentation_Week_9.ppt |Week 9 progress update]] | |||
*[[Media:Cyanobacteria_presentation_Week_10.ppt |Week 10 progress update, 50% complete]] | |||
*''[[Media:Cyanobacteria_final_presentation.ppt |Final Presentation (incomplete)]]'' --old | |||
*''[[Media:final_presentation_draft2.ppt |Final Presentation (complete)]]'' --old | |||
*[[:Image:Cyano presentation.ppt | Jamboree presentation]] (in progress) | |||
**[[:Image:Cyano_presentation_script.doc|Script]] (in progress) | |||
*[[:Image:Cyano poster.ppt | Cyano poster]] (in progress) | |||
== | ==Team Members== | ||
*[[User:Hetmann|Hetmann Hsieh]] ([[User_talk:Hetmann|talk]], [[Special:Contributions/Hetmann|edits]]) | |||
* | *[[User:JeffreyLau|Jeffrey Lau]] ([[User_talk:JeffreyLau|talk]], [[Special:Contributions/JeffreyLau|edits]]) | ||
* | *[[User:Zhipeng Sun|Zhipeng Sun]] ([[User_talk:Zhipeng_Sun|talk]], [[Special:Contributions/Zhipeng_Sun|edits]]) | ||
*[[User:DavidRamos|David Ramos]] ([[User_talk:DavidRamos|talk]], [[Special:Contributions/DavidRamos|edits]]) | |||
== | ==Recent Changes== | ||
{{Special:Recentchanges/b=IGEM:Harvard/2006/Cyanobacteria&limit=25}} | |||
Latest revision as of 04:28, 3 November 2006
<html><style type='text/css'> .tabs {
font-size:80%;
font-weight:none;
width: 100%;
color: #FFFFFF;
background:#FFFFFF url("/images/5/54/DarkgreenTab-bg.gif") repeat-x bottom;
}
.tabs li {
background:url("/images/3/36/DarkgeenTab-left.gif") no-repeat left top;
}
.tabs a,.tabs strong {
background:url("/images/d/d3/DarkgreenTab-right.gif") no-repeat right top;
color:#FFFFFF;
padding: 3px 10px 3px 4px;
}
.tabs strong{
color:#CCFF00;
background-image:url("/images/b/b1/DarkgreenTab-right_on.gif");
}
.tabs a:hover{
color:#66FF00;
}
</style></html>
Introduction
Welcome to the lab notebook for the Cyanobacteria oscillator project! The goal of our team, composed of four members, is to reconstruct the cyanobacterial circadian oscillator system into E. coli. Three proteins, KaiA, B, and C, have been shown to have an in-vitro phosphorylation state oscillation (Nakajima et al. 2005) by transcriptional-translational independent methods. If this system can be reconstituted in E. coli, there are two important applications:
- Synthetic Biology: Creating a functional, oscillating set of proteins is the next logical step from the synthetic "repressilator" system engineered by Elowitz et al. (2000). Although a good proof of concept, the "repressilator" lacks the stability needed from a robust oscillator such as the naturally evolved cyanobacterial oscillator. This robust oscillator could prove useful in an eventual biocircuit.
- Circadian Biology: Cyanobacteria are the simplest model organisms for the study of circadian oscillation. Although circadian oscillation has been fairly well characterized, less is understood at the molecular level. By porting the oscillation system into E. coli, one can begin to understand more precisely the pathways involved in the genomic oscillation of cyanobacteria.
For more background information on the ciracadian system, please check out our "Literature" section. Otherwise, day-to-day work can be found under the "Lab Notebook" tab; we will post major results of our work and links to the days as they become available. If you have questions or comments, feel free to contact us: information is located at the main Harvard iGEM 2006 page. Thanks!
Sincerely,
Zhipeng, Hetmann, Dave, and Jeff
Update 10/27/06: We believe we can express the three proteins into e. coli, and that there is interaction between A+C and possible interaction between B+C. See the Lab Notebook for more information.
Outline of Findings and Signifigant Dates
- 07/05/06: The incubator for growing up our cyanobacteria is complete; we have cultures growing! Link
- 07/10/06: Some computer modeling has been done to see the effect of multiple unsyncronized clocks on phosphorylation state output. Link
- 07/21/06: Upon having trouble with site-specific mutagenesis on the KaiA and KaiBC operons from the cyanobacterial genome, we have decided to pursue synthesis of the constructs in parallel with continued extraction attempts. Link
- 08/01/06: Preliminary success with site-specific mutagenesis. Link
- 08/05/06: Promoter leakness tests come out negative. May have to use low-copy plasmids if we want good control of protein expression in Top10F. Link
- 08/11/06: We are moving to the synthetic KaiA, KaiB, and KaiC for future work. Link
- 08/30/06: We successfully made the first construct, Lac+RBS+KaiC. Link
- 09/01/06: Using the newly developed ligation protocol, we have successfully repeated Lac+RBS+KaiC from 08/30/06 and made Lac+RBS+KaiA. Link
- 10/21/06: Successfully made Lac+RBS+KaiB and Lac+RBS+KaiA+Lac+RBS+KaiC. Link
- 10/24/06: Successfully made Lac+RBS+KaiB+Lac+RBS+KaiC. Link
- 10/25/06: Constructs for Stage I have been completed; ready to move to Stage I of Western Blotting, to verify expression of KaiC and interaction of KaiA and KaiB with KaiC. Link
- 10/27/06: Preliminary data indicates that the Kai proteins are being expressed in e. coli and that there is interaction between the three proteins! Link
Construct Planning
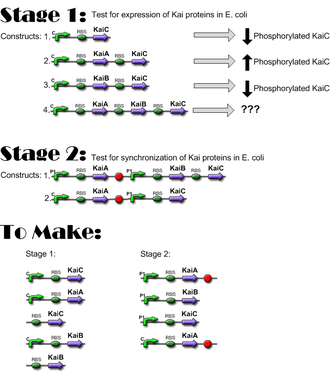
Lengths
From VF2 to VR (BioBrick primers):
- KaiA + J04500: 1406 bp
- KaiB + J04500: 859 bp
- KaiC + J04500: 2110 bp
Agenda
See image at right for our long-term project outline.

BioBricks Used
- <bbpart>BBa_J04450</bbpart>
- RFP device
- Insert size: 1069bp
- [pSB1A2]
- High-copy, AmpR
- Size: 2079bp
- <bbpart>BBa_J04500</bbpart>
- Lac promoter + RBS
- Insert size: 220bp
- [pSB1AK3]
- High-copy, AmpR, KanR
- Insert size: 3189bp
- [pSB4A3]
- Low-copy, AmpR
- Insert size: 3339 bp
- <bbpart>BBa_R0010</bbpart> + <bbpart>BBa_E0241</bbpart>
- GFP device
- Insert size: 995 bp
- <bbpart>BBa_J04450</bbpart>
Presentations
- Project proposal (week 2)
- Week 3 progress update
- Built incubator and obtained WH8102, PCC7942, and PCC6803 strains
- Week 4 progress update
- Week 5 progress update, upd. 10:10 7/17
- Week 6 progress update, upd. 10:02 7/24 HH
- Week 7 progress update
- Week 8 progress update
- Week 9 progress update
- Week 10 progress update, 50% complete
- Final Presentation (incomplete) --old
- Final Presentation (complete) --old
- Jamboree presentation (in progress)
- Script (in progress)
- Cyano poster (in progress)
Team Members
- Hetmann Hsieh (talk, edits)
- Jeffrey Lau (talk, edits)
- Zhipeng Sun (talk, edits)
- David Ramos (talk, edits)
Recent Changes
- N
- This edit created a new page (also see list of new pages)
- m
- This is a minor edit
- b
- This edit was performed by a bot
- (±123)
- The page size changed by this number of bytes
18 April 2024
| 15:01 | Pan:Who we are diffhist +14 Taopan talk contribs | ||||
|
|
15:00 | Pan:Methods 2 changes history +456 [Taopan (2×)] | |||
|
|
15:00 (cur | prev) +2 Taopan talk contribs | ||||
|
|
14:59 (cur | prev) +454 Taopan talk contribs | ||||
|
|
14:56 | Pan:Publications 2 changes history +396 [Taopan (2×)] | |||
|
|
14:56 (cur | prev) +74 Taopan talk contribs | ||||
|
|
14:54 (cur | prev) +322 Taopan talk contribs | ||||
| 13:03 | BioMicroCenter:Pricing diffhist +166 Challee talk contribs | ||||
|
|
12:58 | BioMicroCenter:Singular Sequencing 2 changes history +124 [Challee (2×)] | |||
|
|
12:58 (cur | prev) +14 Challee talk contribs (→Things to Consider) | ||||
|
|
12:57 (cur | prev) +110 Challee talk contribs | ||||
|
|
12:12 | BioMicroCenter:Tecan Freedom Evo 7 changes history +1,746 [Noelani Kamelamela (7×)] | |||
|
|
12:12 (cur | prev) +4 Noelani Kamelamela talk contribs | ||||
|
|
12:12 (cur | prev) +3 Noelani Kamelamela talk contribs | ||||
|
|
10:13 (cur | prev) +7 Noelani Kamelamela talk contribs (→verrity Chemagic 360) | ||||
|
|
10:08 (cur | prev) −42 Noelani Kamelamela talk contribs (→verrity Chemagic 360) | ||||
|
|
10:08 (cur | prev) +86 Noelani Kamelamela talk contribs (→verrity Chemagic 360) | ||||
|
|
09:34 (cur | prev) +23 Noelani Kamelamela talk contribs (→verrity Chemagic 360) | ||||
|
|
09:32 (cur | prev) +1,665 Noelani Kamelamela talk contribs | ||||
| 11:42 | 3D Cell Culture - McLean Taggart, Emma Villares, Maximillian Marek, Scott LeBlanc, Adam Lyons and Jacob Belden diffhist −3 Sarah L. Perry talk contribs | ||||
|
|
09:35 | BioMicroCenter 2 changes history +92 [Noelani Kamelamela (2×)] | |||
|
|
09:35 (cur | prev) +60 Noelani Kamelamela talk contribs | ||||
|
|
09:20 (cur | prev) +32 Noelani Kamelamela talk contribs | ||||
| 09:32 | Upload log Noelani Kamelamela talk contribs uploaded File:Chemagic360.jpg (from manual) | ||||
17 April 2024
|
|
15:34 | BioMicroCenter:Element Sequencing 3 changes history +295 [Challee (3×)] | |||
|
|
15:34 (cur | prev) +195 Challee talk contribs | ||||
|
|
14:22 (cur | prev) +100 Challee talk contribs | ||||
|
|
14:07 (cur | prev) 0 Challee talk contribs | ||||
| 13:10 | BioMicroCenter:SingleCell diffhist +30 Noelani Kamelamela talk contribs (→10X CHROMIUM X) | ||||
| 12:43 | BioMicroCenter diffhist −15 Noelani Kamelamela talk contribs | ||||
16 April 2024
|
|
N 19:59 | Nanoimprint Lithography (NIL) - Carter Paul 10 changes history +7,205 [CarterPaul (10×)] | |||
|
|
19:59 (cur | prev) +769 CarterPaul talk contribs (→Thermal NIL Process) | ||||
|
|
19:53 (cur | prev) 0 CarterPaul talk contribs (→Thermal NIL Process) | ||||
|
|
19:53 (cur | prev) 0 CarterPaul talk contribs (→Thermal NIL Process) | ||||
|
|
19:52 (cur | prev) +1 CarterPaul talk contribs (→Thermal NIL Process) | ||||
|
|
19:50 (cur | prev) +202 CarterPaul talk contribs (→Thermal NIL Process) | ||||
|
|
19:17 (cur | prev) −20 CarterPaul talk contribs (→References) | ||||
|
|
19:17 (cur | prev) −1 CarterPaul talk contribs | ||||
|
|
19:11 (cur | prev) +4,278 CarterPaul talk contribs | ||||
|
|
18:53 (cur | prev) +1,891 CarterPaul talk contribs | ||||
| N |
|
18:42 (cur | prev) +85 CarterPaul talk contribs (Created page with "{{Template:CHEM-ENG590E}} =Motivation= =Introduction to NIL= =Thermal NIL Process=") | |||
| 19:40 | Upload log CarterPaul talk contribs uploaded File:NIL1.png | ||||
| N 18:40 | 3D Cell Culture - McLean Taggart, Emma Villares, Maximillian Marek, Scott LeBlanc, Adam Lyons and Jacob Belden diffhist +24,060 CarterPaul talk contribs (Created page with "{{Template:CHEM-ENG590E}} ==Introduction== While most microfluidic devices incorporate a 2D cell culture design, in which a single layer of cells is grown on the bottom of a device, these systems suffer from poor <i>in vivo</i> mimicry, as, in the human body, most cells grow in all directions.<sup>https://doi.org/10.5114/aoms.2016.63743 1</sup> To address this limitation, 3D cell culture devices have been developed - in w...") | ||||
|
|
18:38 | CHEM-ENG590E:Wiki Textbook 2 changes history +63 [CarterPaul (2×)] | |||
|
|
18:38 (cur | prev) +50 CarterPaul talk contribs (→Chapter 1 - Microfabrication) | ||||
|
|
18:37 (cur | prev) +13 CarterPaul talk contribs | ||||
| 18:36 | 3D Cell Culture - McLean Taggart, Emma Villares, Maximillian Marek, Scott LeBlanc, and Adam Lyons diffhist +5,343 CarterPaul talk contribs (Added a Technique and applications section) | ||||
|
|
10:20 | Yarn Microfluidics - Roger Dirth 11 changes history +406 [Rcostello (11×)] | |||
|
|
10:20 (cur | prev) +41 Rcostello talk contribs (→Applications) | ||||
|
|
10:19 (cur | prev) +36 Rcostello talk contribs (→Applications) | ||||
|
|
10:18 (cur | prev) +36 Rcostello talk contribs (→Introduction) | ||||
|
|
10:17 (cur | prev) +38 Rcostello talk contribs (→Fabrication) | ||||
|
|
10:17 (cur | prev) +38 Rcostello talk contribs (→Washburn Equation) | ||||
|
|
10:16 (cur | prev) +38 Rcostello talk contribs (→Wicking Rate) | ||||
|
|
10:16 (cur | prev) +37 Rcostello talk contribs (→Introduction) | ||||
|
|
10:15 (cur | prev) +36 Rcostello talk contribs (→Wicking Rate) | ||||
|
|
10:14 (cur | prev) +36 Rcostello talk contribs (→Fabrication) | ||||
|
|
10:14 (cur | prev) +34 Rcostello talk contribs (→Applications) | ||||
|
|
10:14 (cur | prev) +36 Rcostello talk contribs (→Introduction) | ||||
