IGEM:Harvard/2009/Notebook/Harvard iGEM 2010/2010/07/06: Difference between revisions
| Line 148: | Line 148: | ||
===Tagging Miraculin and Brazzein with YFP and StrepII tags=== | ===Tagging Miraculin and Brazzein with YFP and StrepII tags=== | ||
Tag: -E--N--X--<b>STREP/YFP</b>--S--N--P- </ | Tag: -E--N--X--<b>STREP/YFP</b>--S--N--P- <br/> | ||
Insert: E--N--X--<b>Miraculin/Brazzein</b>--S--N--P- </ | Insert: E--N--X--<b>Miraculin/Brazzein</b>--S--N--P- <br/> | ||
*The 'tag' biobrick (either YFP or STREPII) was used as the vector; Miraculin and Brazzein were used as the insert. | *The 'tag' biobrick (either YFP or STREPII) was used as the vector; Miraculin and Brazzein were used as the insert. | ||
Revision as of 14:16, 6 July 2010
 iGEM iGarden iGEM iGarden
|
<html><img src="/images/9/94/Report.png" border="0" /></html> Main project page <html><img src="/images/c/c3/Resultset_previous.png" border="0" /></html>Previous entry<html> </html>Next entry<html><img src="/images/5/5c/Resultset_next.png" border="0" /></html> | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Team AllergyToday, we isolated plasmid that contained antisense and sense sequences of the allergen panel and introns PME and PAL that we grew last week. ProceduresFor amiRNA
For ihpRNA
ResultsFor amiRNAMiniprep of turbo bacteria for plasmids containing sequences for amiRNA Should obtain plasmids containing the complete sequence for amiRNA. The resulting concentration should be relatively high (<100ng/μL) because all growing bacteria would have the plasmids inside. Nanodrop of miniprepped amiRNA plasmids We used 2μL of DNA per nanodrop.
Diagnostic digest of amiRNA plasmids Digested 200~500ng of plasmids with EcoR1 and Spe1. We will then run the digested vectors on an E Gel and look for DNA that is the length of the insert. If such DNA appears on the gel, then we will send it off to be sequenced. E Gel of digested amiRNA  We had to run 21 samples, so we used two gels. The orders were: Gel #1
Gel #2
For ihpRNA
We used 2μL of DNA per nanodrop.
Team FenceInoculated Barstar and NLS colonies from Thursday's ligation. Performed PCR on LacIn (E10) following standard PCR measurements as previously described. Digestion of B21 backbone using Xba1 and Pst1 Total volume per reaction: 50 μL
Digestion successful, larger bands removed for gel extraction. Gel extraction performed according to QIAgen gel extraction protocol. Team FlavourTagging Miraculin and Brazzein with YFP and StrepII tagsTag: -E--N--X--STREP/YFP--S--N--P-
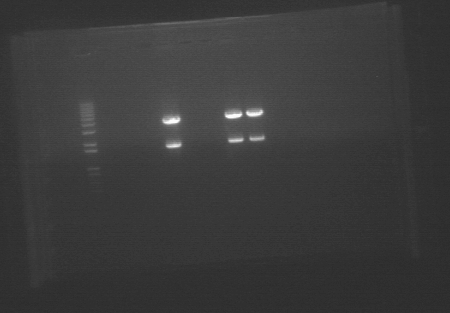
GelThe gel showed DNA fragments consistent with Miraculin and Brazzein. The digestion of B21 appeared to be succesful, but the DNA sequence cut out was too small to see on the gel. Gel Lanes:
1. 1 kb plus ladder
2. B21 speI/pstI
4. B21 ecoRI/xbaI
6. Miraculin xbaI/pstI
8. Miraculin ecoRI/speI
10. Brazzein xbaI/pstI
12. Brazzein ecoRI/pstI
14. 1kb plus ladder
ODs of gel purified DNA
B21 speI/pstI: 15.9 ng/μL
B21 ecoRI/xbaI: 11.0 ng/μL
Miraculin xbaI/pstI: 9.7 ng/μL
Miraculin ecoRI/speI: 7.1 ng/μL
Brazzein xbaI/pstI: 2.6 ng/μL
Brazzein ecoRI/pstI: 5.0 ng/μL
Ligation | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||