IGEM:Paris Bettencourt 2012/Notebooks/suicide group/day by day//2012/09/20: Difference between revisions
From OpenWetWare
Jump to navigationJump to search
| Line 25: | Line 25: | ||
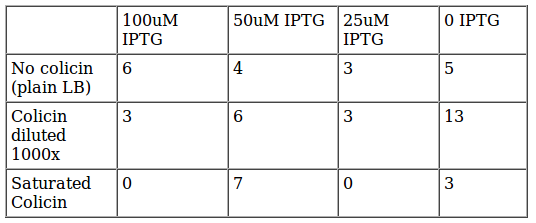
The table shows the number of colonies survive in each plate IPTG x Colicin. Again we had a slighty random result and a very few colonies (theoretically we should get ~200 colonies if all cells survive, and we expected the cells survive if we do not mix them with Colicin), so we decided to repeat the experiment with less dilution. | The table shows the number of colonies survive in each plate IPTG x Colicin. Again we had a slighty random result and a very few colonies (theoretically we should get ~200 colonies if all cells survive, and we expected the cells survive if we do not mix them with Colicin), so we decided to repeat the experiment with less dilution. | ||
[[Image:SG_exp200912.png | [[Image:SG_exp200912.png]] | ||
<!-- ##### DO NOT edit below this line unless you know what you are doing. ##### --> | <!-- ##### DO NOT edit below this line unless you know what you are doing. ##### --> | ||
Revision as of 16:17, 24 September 2012
| <html><img src="/images/9/94/Report.png" border="0" /></html> Main project page <html><img src="/images/c/c3/Resultset_previous.png" border="0" /></html>Previous entry<html> </html>Next entry<html><img src="/images/5/5c/Resultset_next.png" border="0" /></html> | |
pLac + Colicin E2 Immunity CharacterizationWe want to characterize the immunity by varying the concentration of IPTG and the ColicinE2 cells. Protocol
Note: 1ml of cells of OD 1.0 has 1e9 cells, so we expect if all cells survive we will get 200 colonies at the end. ResultsThe table shows the number of colonies survive in each plate IPTG x Colicin. Again we had a slighty random result and a very few colonies (theoretically we should get ~200 colonies if all cells survive, and we expected the cells survive if we do not mix them with Colicin), so we decided to repeat the experiment with less dilution. | |