In-fusion biobrick assembly: Difference between revisions
| Line 25: | Line 25: | ||
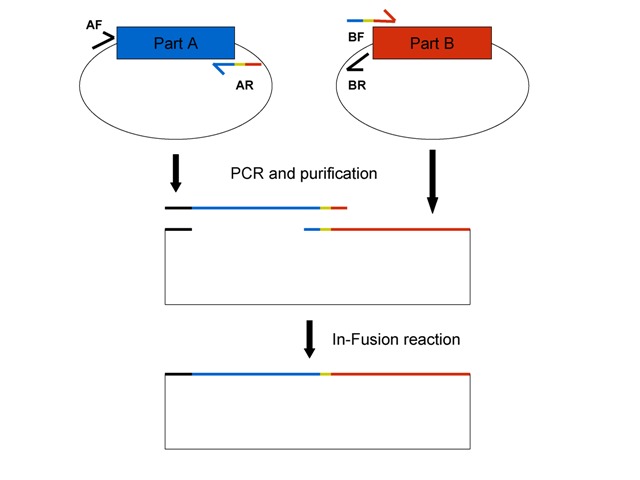
This figure shows the '''general''' In-Fusion BioBrick assembly scheme. See details below. Part A (blue) and Part B (red) are on the same type of plasmid. Primers described below are color-coded to show their homology. The thick black line indicates prefix homology and the yellow sequence is the scar that is normally between parts after standard BioBrick assembly, if this is desired. | This figure shows the '''general''' In-Fusion BioBrick assembly scheme. See details below. Part A (blue) and Part B (red) are on the same type of plasmid. Primers described below are color-coded to show their homology. The thick black line indicates prefix homology and the yellow sequence is the scar that is normally between parts after standard BioBrick assembly, if this is desired. | ||
1. Order primers for parts to assemble. Primers can be optimized accordingly, but this general scheme should work for most BioBricks. The main rule is that the forward primer of one PCR-amplified fragment needs to have at least 15bp homology to the reverse primer of the second fragment, and vice versa, for the In-Fusion reaction to work. | 1. Order primers for parts to assemble. Primers can be optimized accordingly, but this general scheme should work for most BioBricks. The main rule is that the forward primer of one PCR-amplified fragment needs to have at least 15bp homology to the reverse primer of the second fragment, and vice versa, for the In-Fusion reaction to work. It is better to put the vector on the smaller BioBrick since the closer the fragments to fuse are in size, the better the reaction will work, in my experience. | ||
For Part A (upstream part = insert) + Part B with the pSB1A2 vector (downstream part = vector) | For Part A (upstream part = insert) + Part B with the pSB1A2 vector (downstream part = vector) | ||
Revision as of 21:07, 8 April 2009
Overview
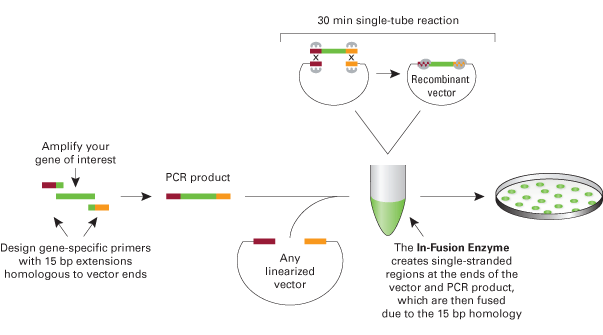
This is a protocol to assemble two BioBricks using the Clontech In-Fusion PCR Cloning Kit and maintains BioBrick standard formats. One PCR-amplified BioBrick has homology on each end with the second PCR-amplified BioBrick (vector amplified with the BioBrick) to allow for the fragments to be fused together in the In-Fusion reaction (see figure below). This procedure can be adapted to fuse more than two fragments together or to re-engineer existing BioBricks. The advantages of In-Fusion BioBrick assembly over standard assembly are that it is faster, does not require restriction digestions or ligations, and is more flexible in the sense that there is more control over the exact engineered sequence. The disadvantages are that it is more expensive, custom primers are required, and occasionally there are mutations in assembled plasmids. However, sequencing a few miniprepped plasmids usually results in at least one plasmid without mutations.
Materials
- Thermocycler
- Primers
- Phusion PCR Mastermix(or another high fidelity polymerase mastermix)
- Qiagen PCR Purification Kit
- Nanodrop (not required, but very useful)
- In-Fusion PCR Cloning Kit
- Gel box, power supply, and gel supplies
- LB+Amp plates
- LB+Amp media
- Qiagen Miniprep Kit
Procedure
This figure shows the general In-Fusion BioBrick assembly scheme. See details below. Part A (blue) and Part B (red) are on the same type of plasmid. Primers described below are color-coded to show their homology. The thick black line indicates prefix homology and the yellow sequence is the scar that is normally between parts after standard BioBrick assembly, if this is desired.
1. Order primers for parts to assemble. Primers can be optimized accordingly, but this general scheme should work for most BioBricks. The main rule is that the forward primer of one PCR-amplified fragment needs to have at least 15bp homology to the reverse primer of the second fragment, and vice versa, for the In-Fusion reaction to work. It is better to put the vector on the smaller BioBrick since the closer the fragments to fuse are in size, the better the reaction will work, in my experience.
For Part A (upstream part = insert) + Part B with the pSB1A2 vector (downstream part = vector)
- Part A Forward primer (AF): 5'-TTCTGGAATTCGCGGCCGCTTCTAG-3' (specific to the pSB1A2 prefix + 5 bases upstream of the prefix)
- Part A Reverse primer (AR): last 20 bases Part A + scar (if wanted) + first 20 bases Part B (reverse complement)
- Part B + vector Forward primer (BF): reverse complement of Part A reverse primer
- Part B + vector reverse primer (BR): 5'-CTAGAAGCGGCCGCGAATTCCAGAA-3' (reverse complement of Part A forward primer)
For Part A with the vector + Part B
- Part A + vector Forward primer: 5'-TACTAGTAGCGGCCGCTGCAGGCTTC-3' (specific to the pSB1A2 suffix + 5 bases downstream of the prefix)
- Part A + vector Reverse primer: last 20 bases Part A + scar (if wanted) + first 20 bases Part B (reverse complement)
- Part B Forward primer: reverse complement of Part A reverse primer
- Part B reverse primer: 5'-GAAGCCTGCAGCGGCCGCTACTAGTA-3' (reverse complement of Part A forward primer)
2. Perform a PCR reaction with a high fidelity polymerase such as Phusion using the recommended protocol. For template DNA, dilute miniprepped BioBrick plasmid DNA 1:1000 to minimize these “background” plasmids from getting transformed later. Digestion with DpnI after the PCR reaction may reduce background plasmids, but in my experience there is a high success rate (>50%) without the use of DpnI or phosphatase on the vector. Using the Cloning Enhancer on the insert provided in some In-Fusion kits gives a much higher number of transformants in my experience.
3. Purify PCR products with Qiagen PCR Purification Kit and elute with 30 ul water.
4. Run a gel with 5 ul of the purified PCR products to determine if the products are of the correct size and there are no secondary bands.
5. Measure the concentration of your PCR products with a Nanodrop or by comparing against a ladder on your gel.
6. Perform the In-Fusion reaction using the recommended protocol (this includes the transformation step).
7. Perform colony PCR on at least 3 colonies (10 is ideal) using the VF2/VR primers to find colonies that have the correct size insert.
8. Grow correct colony or colonies in LB+Amp overnight.
9. Perform miniprep(s) with the Qiagen Miniprep Kit.
10. Submit plasmid(s) for sequencing.
Notes
Please feel free to post comments, questions, or improvements to this protocol. Happy to have your input!
- List troubleshooting tips here.
- You can also link to FAQs/tips provided by other sources such as the manufacturer or other websites.
- Anecdotal observations that might be of use to others can also be posted here.
Please sign your name to your note by adding '''*~~~~''': to the beginning of your tip.
References
Will enter later.
Contact
sleight at u.washington.edu