Kafatos:Research: Difference between revisions
No edit summary |
No edit summary |
||
| (2 intermediate revisions by the same user not shown) | |||
| Line 77: | Line 77: | ||
</blockquote> | </blockquote> | ||
|} | |} | ||
{|cellspacing="5" cellpadding="10" style="background:#00538D; width: 800px;" | {|cellspacing="5" cellpadding="10" style="background:#00538D; width: 800px;" | ||
|-valign="top" | |-valign="top" | ||
|style="background:#ffffff"| | |style="background:#ffffff"| | ||
== | ==BBSRC Research Grant on:<BR> Genomic analysis of NF-kappaB signalling in Anopheles gambiae== | ||
<blockquote> | <blockquote> | ||
The innate immune system is the first line of defence against infections in higher organisms. In insects, which like other invertebrate animals lack adaptive immune systems, innate immunity is the only defence system. Innate immunity relies on receptors that recognize specific molecular structures shared between microbes or danger signals generated during an infection. Recognition triggers specific signalling pathways, cellular processes or enzymatic cascades, which activate various effector mechanisms that combat infections. In many cases, the effector mechanisms require de novo production of effector or regulatory proteins, which is usually controlled by transcription factors of the Rel/Nuclear Factor/kappaB (NF-kappaB) family. This project aims to dissect the mechanisms of gene expression that are under the control of NF-kappaB signalling pathways in the African mosquito Anopheles gambiae. This mosquito is a vector for animal and human diseases including malaria that is caused by the protozoan parasite Plasmodium. In recent years, thanks to the availability of its genome sequence and development of robust genetics and genomics tools to investigate the gene function, A. gambiae has become a model system to study the interactions between microbes, especially Plasmodium, and the innate immune system. | The innate immune system is the first line of defence against infections in higher organisms. In insects, which like other invertebrate animals lack adaptive immune systems, innate immunity is the only defence system. Innate immunity relies on receptors that recognize specific molecular structures shared between microbes or danger signals generated during an infection. Recognition triggers specific signalling pathways, cellular processes or enzymatic cascades, which activate various effector mechanisms that combat infections. In many cases, the effector mechanisms require de novo production of effector or regulatory proteins, which is usually controlled by transcription factors of the Rel/Nuclear Factor/kappaB (NF-kappaB) family. This BBSRC-funded project aims to dissect the mechanisms of gene expression that are under the control of NF-kappaB signalling pathways in the African mosquito Anopheles gambiae. This mosquito is a vector for animal and human diseases including malaria that is caused by the protozoan parasite Plasmodium. In recent years, thanks to the availability of its genome sequence and development of robust genetics and genomics tools to investigate the gene function, A. gambiae has become a model system to study the interactions between microbes, especially Plasmodium, and the innate immune system.<BR><BR> | ||
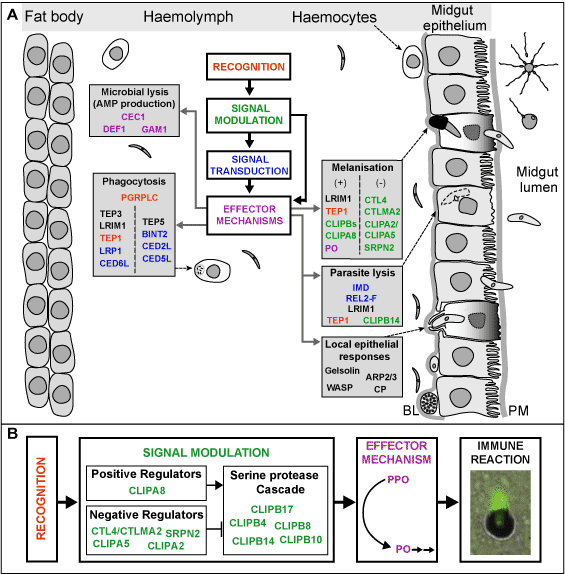
We had previously shown that the mosquito REL2 signalling pathway, which is equivalent to the Drosophila Imd, is responsible for limiting infections of the rodent malaria parasite Plasmodium berghei and that the same pathway is required for resistance against infections with gram-positive and gram-negative bacteria (Meister et al., PNAS, 2005). This project significantly advanced our understanding of the molecular mechanisms that regulate these reactions. We have shown that these two phenomena are tightly linked (Meister et al., PLoS Pathogens, 2009). Infections with bacteria are sensed by the peptidoglycan recognition receptor PGRPLC and activate the REL2 pathway. In this way, the REL2 pathway controls the size of bacteria populations residing in the mosquito midgut, which dramatically increase soon after ingestion of blood. In addition to limiting the bacterial infection, downstream effectors of this pathway also act upon Plasmodium ookinetes invading the mosquito gut, killing a substantial fraction of them. Importantly, this mechanism reduces mosquito infections with field isolates of the deadliest of the human malaria parasites, Plasmodium falciparum, as well as with the laboratory model parasite, Plasmodium berghei. These findings open new research avenues towards understanding the mosquito/parasite interactions, in which symbiotic microbes play a central modulatory role. | We had previously shown that the mosquito REL2 signalling pathway, which is equivalent to the Drosophila Imd, is responsible for limiting infections of the rodent malaria parasite Plasmodium berghei and that the same pathway is required for resistance against infections with gram-positive and gram-negative bacteria (Meister et al., PNAS, 2005). This project significantly advanced our understanding of the molecular mechanisms that regulate these reactions. We have shown that these two phenomena are tightly linked (Meister et al., PLoS Pathogens, 2009). Infections with bacteria are sensed by the peptidoglycan recognition receptor PGRPLC and activate the REL2 pathway. In this way, the REL2 pathway controls the size of bacteria populations residing in the mosquito midgut, which dramatically increase soon after ingestion of blood. In addition to limiting the bacterial infection, downstream effectors of this pathway also act upon Plasmodium ookinetes invading the mosquito gut, killing a substantial fraction of them. Importantly, this mechanism reduces mosquito infections with field isolates of the deadliest of the human malaria parasites, Plasmodium falciparum, as well as with the laboratory model parasite, Plasmodium berghei. These findings open new research avenues towards understanding the mosquito/parasite interactions, in which symbiotic microbes play a central modulatory role.<BR><BR> | ||
Two of the downstream effectors of the mosquito NF-kappaB pathways are the leucine-rich repeat proteins LRIM1 and APL1C. We and others had previously shown that these hemolymph proteins are important antagonists of Plasmodium infections. In the context of this project we have generated substantial new knowledge of the molecular mechanisms implicated in this process (Povelones et al., Science, 2009). We have shown that LRIM1 and APL1C circulate in the mosquito hemolymph as a complex that physically interacts with the complement C3-like protein, TEP1, promoting its cleavage and/or stabilization and localization on the surface of parasites, ultimately causing their lysis or melanization. These data establish a novel mechanism of complement pathway activation in insects. | Two of the downstream effectors of the mosquito NF-kappaB pathways are the leucine-rich repeat proteins LRIM1 and APL1C. We and others had previously shown that these hemolymph proteins are important antagonists of Plasmodium infections. In the context of this project we have generated substantial new knowledge of the molecular mechanisms implicated in this process (Povelones et al., Science, 2009). We have shown that LRIM1 and APL1C circulate in the mosquito hemolymph as a complex that physically interacts with the complement C3-like protein, TEP1, promoting its cleavage and/or stabilization and localization on the surface of parasites, ultimately causing their lysis or melanization. These data establish a novel mechanism of complement pathway activation in insects. | ||
To identify novel regulators of the two NF-kappaB pathways, we have establish high throughput cell-based RNAi screens of over 100 genes expressed in mosquito haemocytes or transcriptionally induced following bacterial infections (Lombardo et al., MPM Proceedings, 2009). We have cloned, tested and used the promoter of the LRIM1 gene, as well as the promoter of the antimicrobial peptide encoding gene CEC1, in luciferase based reporter assays following challenge with various elicitors. Our data have identified several novel regulators of these pathways that are currently being investigated. | To identify novel regulators of the two NF-kappaB pathways, we have establish high throughput cell-based RNAi screens of over 100 genes expressed in mosquito haemocytes or transcriptionally induced following bacterial infections (Lombardo et al., MPM Proceedings, 2009). We have cloned, tested and used the promoter of the LRIM1 gene, as well as the promoter of the antimicrobial peptide encoding gene CEC1, in luciferase based reporter assays following challenge with various elicitors. Our data have identified several novel regulators of these pathways that are currently being investigated.<BR><BR> | ||
Finally, in collaboration with the Broad Institute and the Harvard School of Public Health we have developed a novel high-throughput genome-wide genotyping tool (Muskavitch et al., ASTMH Proceedings, 2008). Development of this SNP chip has been enabled by the discovery of abundant single nucleotide polymorphisms (SNPs) through resequencing of multiple strains by a multicentre consortium. The array is based on a filtered set of 400000 SNP assays, 66000 of which yield perfectly accurate SNP calls. Pooled hybridization results using material from multiple mosquitoes exhibit a very high correlation with averaged hybridization results from individual mosquitoes, enabling the use of a single chip to quantitatively describe genome-wide allele frequencies in a large sample of mosquitoes. We expect that this tool will be widely adopted among vector biologists with interests in mapping of genes underlying important traits in A. gambiae and in analysis of mosquito population structure. | Finally, in collaboration with the Broad Institute and the Harvard School of Public Health we have developed a novel high-throughput genome-wide genotyping tool (Muskavitch et al., ASTMH Proceedings, 2008). Development of this SNP chip has been enabled by the discovery of abundant single nucleotide polymorphisms (SNPs) through resequencing of multiple strains by a multicentre consortium. The array is based on a filtered set of 400000 SNP assays, 66000 of which yield perfectly accurate SNP calls. Pooled hybridization results using material from multiple mosquitoes exhibit a very high correlation with averaged hybridization results from individual mosquitoes, enabling the use of a single chip to quantitatively describe genome-wide allele frequencies in a large sample of mosquitoes. We expect that this tool will be widely adopted among vector biologists with interests in mapping of genes underlying important traits in A. gambiae and in analysis of mosquito population structure. | ||
Latest revision as of 06:53, 25 May 2010
| Click here to visit our NEW WEBSITE |
| The content below is most likely out of date. We also have a new lean and mean openwetware area. |
Functional genomics of mosquito vector/malaria parasite interactions
|
Genomic approaches
|
Targeted approaches
|
BBSRC Research Grant on:
|
Malaria and mosquito population dynamics
|