Kafatos:Research: Difference between revisions
No edit summary |
No edit summary |
||
| Line 1: | Line 1: | ||
{{Kafatos/Christophides Lab}} | {{Kafatos/Christophides Lab}} | ||
__NOTOC__ | |||
{|cellspacing="5" cellpadding="10" style="background:#00538D; width: 800px;" | |||
|-valign="top" | |||
|style="background:#ffffff"| | |||
==Functional genomics of mosquito vector/malaria parasite interactions== | ==Functional genomics of mosquito vector/malaria parasite interactions== | ||
<blockquote> | <blockquote> | ||
| Line 7: | Line 10: | ||
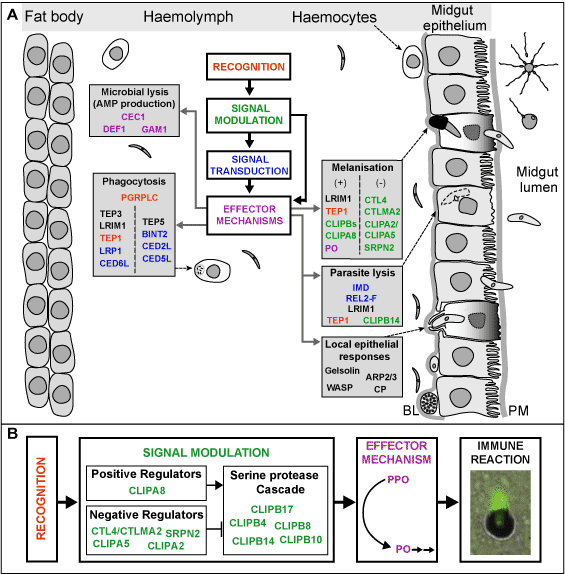
Completion of the genome sequence of ''A. gambiae'' two years ago, together with the development of DNA microarrays in this species and adaptation of the RNAi technique to adult mosquitoes, has allowed comparative and functional genomic approaches to understanding the mosquito innate immune system, and its interactions with parasites. Using the rodent model system, ''P. berghei'', we have identified a variety of factors that negatively affect the development of parasites in the mosquito (antagonists), in some cases leading to complete transmission blockage. In addition, mosquito molecules have been identified that play positive roles and are required for successful parasite transmission (agonists). Research is continuing to identify new factors involved in these interactions and to decipher the interplay of these molecules and their regulation. Importantly, as our findings indicate a highly complex interplay between parasite and vector, we are currently extending our studies of parasite-vector interactions towards the human malaria parasite, ''P. falciparum''. | Completion of the genome sequence of ''A. gambiae'' two years ago, together with the development of DNA microarrays in this species and adaptation of the RNAi technique to adult mosquitoes, has allowed comparative and functional genomic approaches to understanding the mosquito innate immune system, and its interactions with parasites. Using the rodent model system, ''P. berghei'', we have identified a variety of factors that negatively affect the development of parasites in the mosquito (antagonists), in some cases leading to complete transmission blockage. In addition, mosquito molecules have been identified that play positive roles and are required for successful parasite transmission (agonists). Research is continuing to identify new factors involved in these interactions and to decipher the interplay of these molecules and their regulation. Importantly, as our findings indicate a highly complex interplay between parasite and vector, we are currently extending our studies of parasite-vector interactions towards the human malaria parasite, ''P. falciparum''. | ||
</blockquote> | </blockquote> | ||
|} | |||
{|cellspacing="5" cellpadding="10" style="background:#00538D; width: 800px;" | |||
|-valign="top" | |||
|style="background:#ffffff"| | |||
==Genomic approaches== | ==Genomic approaches== | ||
<blockquote> | <blockquote> | ||
| Line 28: | Line 36: | ||
#vlachou2004 pmid=15186403 Real-time, in vivo analysis of malaria ookinete locomotion and mosquito midgut invasion. | #vlachou2004 pmid=15186403 Real-time, in vivo analysis of malaria ookinete locomotion and mosquito midgut invasion. | ||
</biblio> | </biblio> | ||
</blockquote> | |||
|} | |||
{|cellspacing="5" cellpadding="10" style="background:#00538D; width: 800px;" | |||
|-valign="top" | |||
|style="background:#ffffff"| | |||
==Targeted approaches== | ==Targeted approaches== | ||
<blockquote> | <blockquote> | ||
| Line 62: | Line 75: | ||
#ligoxygakis2002 pmid=12456640 | #ligoxygakis2002 pmid=12456640 | ||
</biblio> | </biblio> | ||
</blockquote> | |||
|} | |||
{|cellspacing="5" cellpadding="10" style="background:#00538D; width: 800px;" | |||
|-valign="top" | |||
|style="background:#ffffff"| | |||
==Malaria and mosquito population dynamics== | ==Malaria and mosquito population dynamics== | ||
<blockquote> | <blockquote> | ||
Transmission of the ''Plasmodium'' parasite to a human host completely depends on the availability of a competent mosquito vector. ''A. gambiae'' is the most important and efficient vector for transmission of human malaria in Africa. In some cases, ''A. gambiae'' kills the ''Plasmodium'' parasite, thus blocking the transmission cycle. The interactions between the vector and the parasite involve the mosquito’s innate immunity. Sequence polymorphisms occurring in immune-related genes (as already documented for ''TEP1'') may reflect phenotypic variations in vector competence. Moreover polymorphism may be indicative of adaptation in a co-evolving mosquito/parasite system. We therefore screened four strains of ''A. gambiae'' (susceptible and refractory to ''Plasmodium'' parasites) and wild populations for their polymorphism. Sixty immune-related genes were sequenced, and Single Nucleotide Polymorphisms (SNPs) were identified by alignment. The diversity in immune-related genes was high compared to other parts of the genome, suggesting a diversifying selection acting on these sequences. The identified SNPs will be used to investigate the potential association between genotypes and the susceptibility/refractoriness of ''A. gambiae'' to the parasite under field conditions. | Transmission of the ''Plasmodium'' parasite to a human host completely depends on the availability of a competent mosquito vector. ''A. gambiae'' is the most important and efficient vector for transmission of human malaria in Africa. In some cases, ''A. gambiae'' kills the ''Plasmodium'' parasite, thus blocking the transmission cycle. The interactions between the vector and the parasite involve the mosquito’s innate immunity. Sequence polymorphisms occurring in immune-related genes (as already documented for ''TEP1'') may reflect phenotypic variations in vector competence. Moreover polymorphism may be indicative of adaptation in a co-evolving mosquito/parasite system. We therefore screened four strains of ''A. gambiae'' (susceptible and refractory to ''Plasmodium'' parasites) and wild populations for their polymorphism. Sixty immune-related genes were sequenced, and Single Nucleotide Polymorphisms (SNPs) were identified by alignment. The diversity in immune-related genes was high compared to other parts of the genome, suggesting a diversifying selection acting on these sequences. The identified SNPs will be used to investigate the potential association between genotypes and the susceptibility/refractoriness of ''A. gambiae'' to the parasite under field conditions. | ||
</blockquote> | </blockquote> | ||
|} | |||
Revision as of 04:27, 24 August 2006
| Click here to visit our NEW WEBSITE |
| The content below is most likely out of date. We also have a new lean and mean openwetware area. |
Functional genomics of mosquito vector/malaria parasite interactions
|
Genomic approaches
|
Targeted approaches
|
Malaria and mosquito population dynamics
|