Lee:Research: Difference between revisions
From OpenWetWare
Jump to navigationJump to search
No edit summary |
|||
| (14 intermediate revisions by the same user not shown) | |||
| Line 13: | Line 13: | ||
The production of distinct mRNA transcripts from a single gene via alternative splicing is a common, yet important mechanism of generating proteomic diversity in eukaryotic cells. Alterations in splicing patterns have been increasingly associated with disease development and variable response to drug therapy. We have been investigating the functional significance and regulatory mechanisms of splicing variants of genes encoding membrane transporters or molecular targets for cancer therapy such as proteasomes. | The production of distinct mRNA transcripts from a single gene via alternative splicing is a common, yet important mechanism of generating proteomic diversity in eukaryotic cells. Alterations in splicing patterns have been increasingly associated with disease development and variable response to drug therapy. We have been investigating the functional significance and regulatory mechanisms of splicing variants of genes encoding membrane transporters or molecular targets for cancer therapy such as proteasomes. | ||
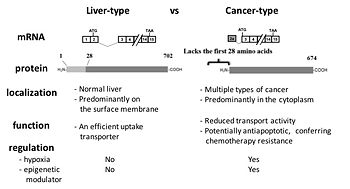
Our group was the first to report that colon and pancreatic cancer cells express the cancer-type variant of OATP1B3 (ct-OATP1B3) with the distinct mRNA and protein identity from the liver-type OATP1B3 detected in non-malignant hepatocytes (Thakkar et al. Mol Pharm 2013). In our follow-up investigation, we identified hypoxia-inducible factor 1alpha (HIF-1alpha) as a positive regulator of ct-OATP1B3 expression (Han et al. Biochem Pharmacol 2013). Recently, we reported that the N-terminal region of OATP1B3 (which is present in liver-type OATP1B3, but absent in ct-OATP1B3) plays a crucial role in regulating membrane trafficking of lt-OATP1B3 (Chun et al. Biochem | Our group was the first to report that colon and pancreatic cancer cells express the cancer-type variant of OATP1B3 (ct-OATP1B3) with the distinct mRNA and protein identity from the liver-type OATP1B3 detected in non-malignant hepatocytes (Thakkar et al. Mol Pharm 2013). In our follow-up investigation, we identified hypoxia-inducible factor 1alpha (HIF-1alpha) as a positive regulator of ct-OATP1B3 expression (Han et al. Biochem Pharmacol 2013). Recently, we reported that the N-terminal region of OATP1B3 (which is present in liver-type OATP1B3, but absent in ct-OATP1B3) plays a crucial role in regulating membrane trafficking of lt-OATP1B3 (Chun et al. Biochem Pharmacol 2017). | ||
We have also been investigating the impact of various genetic variations associated with different subunits that constitute the multi-subunit protease complex called proteasomes. The β1i subunit of the immunoproteasome harbors frequently occurring genetic variations (p.60R>H) and conflicting results had been reported with regard to its functional impact on the immunoproteasome activity. We reported that the codon 60 genetic variations of the β1i subunit do not account for variable expression/activity of the immunoproteasome (Park et al. PLoS One 2013). Currently we are investigating the impact of alternatively spliced transcripts of the proteasome subunits on the proteasome function. | We have also been investigating the impact of various genetic variations associated with different subunits that constitute the multi-subunit protease complex called proteasomes. The β1i subunit of the immunoproteasome harbors frequently occurring genetic variations (p.60R>H) and conflicting results had been reported with regard to its functional impact on the immunoproteasome activity. We reported that the codon 60 genetic variations of the β1i subunit do not account for variable expression/activity of the immunoproteasome (Park et al. PLoS One 2013). Currently, we are investigating the impact of alternatively spliced transcripts of the proteasome subunits on the proteasome function. | ||
# | <biblio> | ||
# | #Thakkar2013 pmid=23215050 (2013) | ||
# | #Han2013 pmid=23924606 (2013) | ||
# | #Park2013 pmid=24040045 (2013) | ||
# | #Thakkar2015 pmid=25735612 (2015) | ||
# | #Chun2017 pmid=28216016 (2017 May) | ||
#Park2017 pmid=28971357 (2017) | |||
#Lee2020 pmid=33051208 (2020) | |||
</biblio> | |||
===II. Development of novel proteasome inhibitor drugs and delivery strategies to improve anticancer efficacy and expand therapeutic utilities=== | ===II. Development of novel proteasome inhibitor drugs and delivery strategies to improve anticancer efficacy and expand therapeutic utilities=== | ||
{|style="background:#ffffff"| | {|style="background:#ffffff"| | ||
[[Image:PI drugs.jpg|thumb|right|450px|FDA-approved proteasome inhibitor drugs]] | [[Image:PI drugs.jpg|thumb|right|450px|FDA-approved proteasome inhibitor drugs]] | ||
The proteasome is | The proteasome is a valid anticancer target, firmly validated by the three FDA-approved drugs, bortezomib (Velcade®), carfilzomib (Kyprolis®) and ixazomib (Ninlalo®). These drugs have transformed the treatment landscape for multiple myeloma and other hematological malignancies. In particular, the second-generation proteasome inhibitor drug carfilzomib has placed itself as part of the frontline multiple myeloma therapy, thanks to its much-improved safety and efficacy profiles over bortezomib. Despite these remarkable successes, carfilzomib has shown limited efficacy in patients with solid cancers, possibly due to its short half-life in blood and insufficient access of active drug to tumor tissues. Working together with experts in the field of drug delivery (Drs. Yoonsoo Bae, University of Kentucky; Dr. Yoon Yeo, Purdue University), we are developing novel nano-formulations for carfilzomib that can improve the biopharmaceutical properties and the anticancer efficacy in multiple myeloma and other types of cancers (Ao et al. J Pharm Exp Ther 2015; Park et al. PLoS One, 2017; Park et al. J Controlled Rel, 2019). | ||
We also collaborate with scientists who have | We also collaborate with scientists who have expertise and experience in the development of novel proteasome inhibitors (Dr. Kyung Bo Kim, University of Kentucky). Our goal is to develop novel proteasome inhibitor drugs that can overcome the biopharmaceutical limitations of existing proteasome inhibitor drugs. Like many cancer therapeutics, proteasome inhibitor drugs are subject to de novo or acquired resistance, which is a major clinical obstacle. We have been investigating the molecular mechanisms underlying cancer resistance to carfilzomib and potential strategies to overcome resistance (Ao et al. Mol. Pharmaceutics 2012). A better mechanistic understanding of cancer resistance to proteasome inhibitors has been utilized to screen and develop novel next-generation proteasome inhibitor drugs that can be effective in patients who do not have any further therapeutic options (Miller et al. J Med Chem 2015; Lee et al. J Med Chem 2019). We also investigate the biological impact and therapeutic potential of novel inhibitors for the immunoproteasome (an alternative type of proteasome that is often upregulated in cancer cells). | ||
# | <biblio> | ||
# | #Ao2015 pmid=26311812 (2015) | ||
# | #Miller2015 pmid=25658656 (2015) | ||
# | #Park2017 pmid=28273121 (2017 March) | ||
# | #Park2018 pmid=29654740 (2018 Aug) | ||
#Park2019 pmid=30954620 (2019 May) | |||
#Lee2019 pmid=30964987 (2019 May) | |||
#Jun2020 pmid=31945310 (2020 April) | |||
</biblio> | |||
===III. Clinical Pharmacokinetics, Pharmacogenomics & Pharmacometrics=== | ===III. Clinical Pharmacokinetics, Pharmacogenomics & Pharmacometrics=== | ||
Interindividual differences in drug response and toxicities are consistently observed with most chemotherapeutic agents or regimens and many clinical variables (e.g., age, gender, diet, drug-drug interactions) affect drug responses. In particular, inherited variations in drug disposition (metabolism and transport) and drug target genes are known to substantially contribute to the observed variability in cancer treatment | Interindividual differences in drug response and toxicities are consistently observed with most chemotherapeutic agents or regimens and many clinical variables (e.g., age, gender, diet, drug-drug interactions) affect drug responses. In particular, inherited variations in drug disposition (metabolism and transport) and drug target genes are known to substantially contribute to the observed variability in cancer treatment outcomes. In our collaborative Phase II clinical study, we investigated the clinical utility of pharmacogenomically selected treatment using genetic polymorphisms in patients with gastric and gastroesophageal junction (GEJ) cancer (Goff et al. PLoS One 2014). Working together with clinical investigators, we investigate the pharmacokinetics of novel cancer drugs, drug formulations or new combination regimens in early clinical trial settings. Our group is also interested in pharmacometric modeling and simulation using preclinical and clinical data. | ||
# | <biblio> | ||
# | #Goff2014 pmid=25232828 (2014) | ||
# | #Wang-Gillam2014 pmid=24916546 (2014) | ||
#Michel2016 pmid=27311401 (2016) | |||
#Asaumi2018 pmid=29368402 (2018) | |||
#Sato2018 pmid=29475833 (2018) | |||
#Nakamura2018 pmid=29920987 (2018 July) | |||
#Nishiyama2019 pmid=30821133 (2019 June) | |||
</biblio> | |||