Maloof Lab:Jose M. Jimenez-Gomez: Difference between revisions
mNo edit summary |
No edit summary |
||
| (7 intermediate revisions by 2 users not shown) | |||
| Line 7: | Line 7: | ||
<h2>Jose M Jimenez-Gomez, PhD.</h2> | <h2>Jose M Jimenez-Gomez, PhD.</h2> | ||
[mailto:jmjimenez@ucdavis.edu Contact] | [mailto:jmjimenez@ucdavis.edu Contact] | ||
<p> | |||
<br> | |||
<font color='red'>The information contained in this website may be outdated.</font><br> | |||
Please use my this website instead: [http://jimenez-gomez_lab.openwetware.org Jimenez Gomez Lab]. | |||
<br> | |||
</p> | |||
|[[Image:Pepe_b&w.jpg|right|135px]] | |[[Image:Pepe_b&w.jpg|right|135px]] | ||
|} | |} | ||
<br> | <br> | ||
| Line 36: | Line 43: | ||
<br> | <br> | ||
<br> | <br> | ||
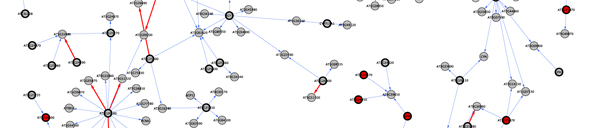
We focused in a chromosomal region containing close to 400 genes to fine map and identify the gene responsible for the differences found in the response to shade in the Bay-0 x Sha population. To do this we employed traditional genetic approaches as well as genomic and network analysis. This network analysis is based on coexpression of the candidate genes with other genes across microarray experiments <cite>Riken</cite>, colocalization with expression QTLs <cite>West07</cite>, functional categorization <cite>GO_Classification</cite> and presence of polymorphisms between the parental lines <cite>Clark07</cite>. The use of this bioinformatic approach allowed us to identify ELF3 as the candidate gene for the shade avoidance QTL, which was then confirmed by traditional | We focused in a chromosomal region containing close to 400 genes to fine map and identify the gene responsible for the differences found in the response to shade in the Bay-0 x Sha population. To do this we employed traditional genetic approaches as well as genomic and network analysis. This network analysis is based on coexpression of the candidate genes with other genes across microarray experiments <cite>Riken</cite>, colocalization with expression QTLs <cite>West07</cite>, functional categorization <cite>GO_Classification</cite> and presence of polymorphisms between the parental lines <cite>Clark07</cite>. The use of this bioinformatic approach allowed us to identify ELF3 as the candidate gene for the shade avoidance QTL, which was then confirmed by traditional fine mapping and cloning. | ||
<br> | <br> | ||
<br> | <br> | ||
| Line 47: | Line 54: | ||
<br> | <br> | ||
<br> | <br> | ||
In the publication of this work, ELF3 alleles of Bay-0 and Sha are shown to differentially affect the shade avoidance response in flowering time and circadian rhtyhms <cite>Jimenez-Gomez10</cite> | |||
<br> | <br> | ||
<br> | <br> | ||
| Line 53: | Line 60: | ||
---- | ---- | ||
<br> | <br> | ||
Tomato is a specially interesting species because of its natural history, phenotypic | Tomato is a specially interesting species because of its natural history, phenotypic diversity among its wild relatives and economic importance. To study the genomic variation among the wild tomato species, we first mined the numerous tomato EST sequences available in the databases in search of polymorphisms. In this dataset, we estimated divergence rates among genes from selected species, and obtained a new set of molecular markers useful in natural variation studies. We performed functional and evolutionary pre-genomic analyses, which gave us an idea of which gene families evolve more rapidly/slowly and have been important during tomato domestication. The results from this work were published <cite>Jimenez-Gomez09</cite> and are available to the community [http://www.plb.ucdavis.edu/labs/maloof/TomatoSNP/index.asp here]. | ||
Now, we are using RNAseq to sequence the transcriptome of four tomato species grown in sun and shade: <i>S. lycopersicum var M82</i>, <i>S. pennellii</i>, <i>S. pimpinellifollium</i> and <i>S. habrochaites</i>. | Now, we are using RNAseq to sequence the transcriptome of four tomato species grown in sun and shade: <i>S. lycopersicum var M82</i>, <i>S. pennellii</i>, <i>S. pimpinellifollium</i> and <i>S. habrochaites</i>. | ||
We developed bioinformatic pipelines to analyze the more than 400 million reads obtained fronm different tissues, species and conditions. The pipeline include scripts that filter and map the reads, detect polymorphisms, calculte their effect on the proteins, perform evolutionary analyses and calculate genome-wide expression levels. Using this methods we | We developed bioinformatic pipelines to analyze the more than 400 million reads obtained fronm different tissues, species and conditions. The pipeline include scripts that filter and map the reads, detect polymorphisms, calculte their effect on the proteins, perform evolutionary analyses and calculate genome-wide expression levels. Using this methods we identified more than 500.000 polymorphisms in these four speceies and calculated expression differences between species, tissues and environmental conditions. | ||
<br> | <br> | ||
| Line 84: | Line 91: | ||
#GO_Classification pmid=10802651 | #GO_Classification pmid=10802651 | ||
#Jimenez-Gomez09 pmid=19575805 | #Jimenez-Gomez09 pmid=19575805 | ||
#Jimenez-Gomez10 pmid=20838594 | |||
</biblio> | </biblio> | ||
|} | |} | ||
__NOTOC__ | __NOTOC__ | ||
Latest revision as of 05:29, 11 January 2012
|
Room 2115 |
QTL and Network analysis of the shade avoidance response in Arabidopsis
Expresion profiling and Single Nuncleotide Polymorphism discovery in cultivated tomato and its wild relatives
Molecular evolution of PHYTOCHROME B
References
|