Maloof Lab:Jose M. Jimenez-Gomez: Difference between revisions
No edit summary |
No edit summary |
||
| Line 28: | Line 28: | ||
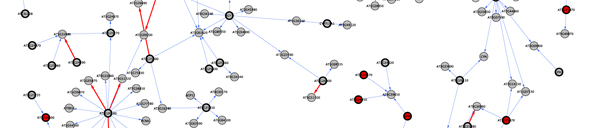
We are focusing now in a chromosomal region containing about 200 genes to fine map and identify the gene responsible for the differential response to shade between the two natural populations. To do this we employ traditional genetic approaches as well as genomic and network analysis. We are developing a protocol to construct gene networks that will help us consider candidate genes based on coexpression with other genes across all microarray experiments performed in Arabidopsis, colocalization with expression QTLs (West at el. 07), functional categorization and presence of polymorphisms (Clark et al. 07). | We are focusing now in a chromosomal region containing about 200 genes to fine map and identify the gene responsible for the differential response to shade between the two natural populations. To do this we employ traditional genetic approaches as well as genomic and network analysis. We are developing a protocol to construct gene networks that will help us consider candidate genes based on coexpression with other genes across all microarray experiments performed in Arabidopsis, colocalization with expression QTLs (West at el. 07), functional categorization and presence of polymorphisms (Clark et al. 07). | ||
[[Image: | [[Image:Network_fragment.jpg]] | ||
Revision as of 22:12, 23 November 2007
|
Room 2115 |
QTL analysis of the shade avoidance response in ArabidopsisPlants exhibit phenotypic plasticity in response to different environmental light cues. For example, shade from neighboring plants sensed by the phytochrome photoreceptors causes increased petiole and stem elongation and early reproduction, collectively called the Shade Avoidance Response. Interestingly, the degree of plasticity varies among strains and species, and this variation can have adaptive value. Plants form different environments exhibit different degrees of responsiveness to the same light stimulus. For example, when plants accommodated to sunny environments detect foliar shade from neighboring vegetation they respond increasing petiole and stem elongation and reducing the time to reproduction, a phenomenon called the "shade avoidance response". On the other hand, plants surrounded by tall vegetation, used to the shade and do not present this response.
To identify the molecular mechanisms underlying this differences we are performing QTL analysis using a previously developed, well characterized Recombinant Inbred Line set descent from two different natural populations of Arabidopsis thaliana: Bayreuth, originary from the German low altitude fallow lands, and Shahdara, from the high mountains of Tadjikistan (Loudet et al. 02). We are focusing now in a chromosomal region containing about 200 genes to fine map and identify the gene responsible for the differential response to shade between the two natural populations. To do this we employ traditional genetic approaches as well as genomic and network analysis. We are developing a protocol to construct gene networks that will help us consider candidate genes based on coexpression with other genes across all microarray experiments performed in Arabidopsis, colocalization with expression QTLs (West at el. 07), functional categorization and presence of polymorphisms (Clark et al. 07).
A more detailed explanation on plant adaptation, responses to light and molecular evolution can be found at the Maloof lab webpage.
|