Quint Lab:Research: Difference between revisions
Marcel Quint (talk | contribs) No edit summary |
Marcel Quint (talk | contribs) |
||
| (4 intermediate revisions by the same user not shown) | |||
| Line 16: | Line 16: | ||
our lab is primarily interested in understanding the genetics and molecular biology of [http://en.wikipedia.org/wiki/Auxin auxin] and other [http://en.wikipedia.org/wiki/Plant_hormone plant hormone] responses in the tiny weed [http://en.wikipedia.org/wiki/Arabidopsis_thaliana arabidopsis thaliana] and related [http://en.wikipedia.org/wiki/Brassicaceae brassicaceae]. phytohormones are one of the classic fields in [http://en.wikipedia.org/wiki/Plant_physiology plant physiology] and the past has shown that understanding hormone action in plants bears great potential for agricultural and horticultural applications. by contributing to the current state of knowledge of hormone biology we hope to participate in the advancement of [http://en.wikipedia.org/wiki/Crop_science crop science]. | our lab is primarily interested in understanding the genetics and molecular biology of [http://en.wikipedia.org/wiki/Auxin auxin] and other [http://en.wikipedia.org/wiki/Plant_hormone plant hormone] responses in the tiny weed [http://en.wikipedia.org/wiki/Arabidopsis_thaliana arabidopsis thaliana] and related [http://en.wikipedia.org/wiki/Brassicaceae brassicaceae]. phytohormones are one of the classic fields in [http://en.wikipedia.org/wiki/Plant_physiology plant physiology] and the past has shown that understanding hormone action in plants bears great potential for agricultural and horticultural applications. by contributing to the current state of knowledge of hormone biology we hope to participate in the advancement of [http://en.wikipedia.org/wiki/Crop_science crop science]. | ||
Secondly, we are fascinated by the mechanisms of [http://en.wikipedia.org/wiki/Molecular_evolution molecular evolution] and how they shape plant | Secondly, we are fascinated by the mechanisms of [http://en.wikipedia.org/wiki/Molecular_evolution molecular evolution] and how they shape plant growth and development. That ranges from learning about the evolutionary history of signaling genes to phylotranscriptomic approaches to plant [http://en.wikipedia.org/wiki/Evo_devoevo evo devo] research. | ||
<br> | <br> | ||
we apply mostly [http://en.wikipedia.org/wiki/Genomics genomics] approaches, such as: | we apply mostly [http://en.wikipedia.org/wiki/Genomics genomics] approaches, such as: | ||
| Line 53: | Line 53: | ||
|width=790px style="padding: 5px; background-color: #ffffff; border: 2px solid #C9D3EB;" | | |width=790px style="padding: 5px; background-color: #ffffff; border: 2px solid #C9D3EB;" | | ||
===evo | ===evo devo=== | ||
[[Image:Hourglass_mosaic.jpg|100px|right]] | [[Image:Hourglass_mosaic.jpg|100px|right]] | ||
for one, we are interested in the evolutionary history of gene families that are involved in important signaling cascades, such as the ubiquitin-proteasome system (see schumann et al. [http://www.ncbi.nlm.nih.gov/pubmed/21119043 ''plant physiology 2011'']). furthermore, we are developing ways to utilize whole genome transcriptional information for evolutionary approaches in close collaboration with the lab of [http://www.informatik.uni-halle.de/arbeitsgruppen/bioinformatik/mitarbeiterinnen/grosse/?lang=en ivo grosse]. by applying phylotranscriptomics – the combination of [http://en.wikipedia.org/wiki/Phylogenetics phylogenetics] and [http://en.wikipedia.org/wiki/Transcriptomics transcriptomics] – to developmental series such as embryogenesis, we are able to trace the evolutionary path across a complete developmental process. | for one, we are interested in the evolutionary history of gene families that are involved in important signaling cascades, such as the ubiquitin-proteasome system (see schumann et al., [http://www.ncbi.nlm.nih.gov/pubmed/21119043 ''plant physiology 2011'']). furthermore, we are developing ways to utilize whole genome transcriptional information for evolutionary approaches in close collaboration with the lab of [http://www.informatik.uni-halle.de/arbeitsgruppen/bioinformatik/mitarbeiterinnen/grosse/?lang=en ivo grosse]. by applying phylotranscriptomics – the combination of [http://en.wikipedia.org/wiki/Phylogenetics phylogenetics] and [http://en.wikipedia.org/wiki/Transcriptomics transcriptomics] – to developmental series such as embryogenesis ( see quint et al., [http://www.ncbi.nlm.nih.gov/pubmed/22951968 ''nature 2012'']), we are able to trace the evolutionary path across a complete developmental process. | ||
|} | |} | ||
Latest revision as of 15:53, 13 February 2013
let's play"real science has the potential to not only amaze, but also transform the way one thinks of the world and oneself. this is because the process of science is little different from the deeply resonant, natural processes of play. play enables humans (and other mammals) to discover (and create) relationships and patterns. when one adds rules to play, a game is created. this is science: the process of playing with rules that enables one to reveal previously unseen patterns of relationships that extend our collective understanding of nature." - Blackawton et al., 2011, Biology letters |
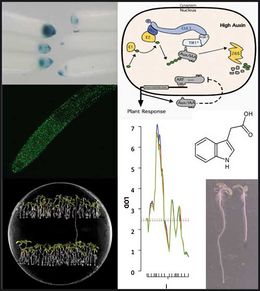 broad research scopeHOW do organisms adapt to the environment and how do they react to different internal and external stimuli? Secondly, we are fascinated by the mechanisms of molecular evolution and how they shape plant growth and development. That ranges from learning about the evolutionary history of signaling genes to phylotranscriptomic approaches to plant evo devo research.
|
 natural variation and quantitative genetics of hormone responsesquantitative genetics

|
evo devo for one, we are interested in the evolutionary history of gene families that are involved in important signaling cascades, such as the ubiquitin-proteasome system (see schumann et al., plant physiology 2011). furthermore, we are developing ways to utilize whole genome transcriptional information for evolutionary approaches in close collaboration with the lab of ivo grosse. by applying phylotranscriptomics – the combination of phylogenetics and transcriptomics – to developmental series such as embryogenesis ( see quint et al., nature 2012), we are able to trace the evolutionary path across a complete developmental process. |
