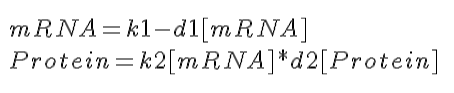User:James Chappell /R RK4 Modeling: Difference between revisions
mNo edit summary |
mNo edit summary |
||
| Line 23: | Line 23: | ||
[[Image:Jec con exp equation.gif]] | |||
library(odesolve) | library(odesolve) | ||
Revision as of 08:56, 8 July 2009
<html> <style type="text/css"> .firstHeading {display: none;} </style> </html>
James Chappell
Imperial College London Synthetic Biology Lab
RK4
The RK4 tool is a tool that can be used to model ordinary differential equations (ODE). Like other ODE solvers you can output the results of a simulation into graph or tables.
Example 1 : Simple Constitutive Expression
library(odesolve)
#recalls the rk4 allgorithm - need to have instal packages to be able to recall and use this function
rm(list=objects())
#Function removes any previous objects from previous simulations using this or other functions, means always need to define any objects for each simulation
(constitutiveexpression <- function(t, x, parms) {
#Define the name of the model, in this case "constitutiveexpression"
mRNA <- x[1]
protein <- x[2]
#Name the response variables – these are the outputs of our model
with(as.list(parms),{
dmRNA <- k1 - d1*mRNA
dprotein <- k2*mRNA - d2*protein
res<-c(dmRNA, dprotein)
list(res)
})}
#Combine these vectors into a list called res
times <-seq(0, 1000, length=1001)
parms <-c(k1=0.69 , k2=f ,d1=0.23 ,d2=0.012)
#Generate a time series over which to solve the equations (1000 in steps of 1)
#Set the parameter values in parms
y <- xstart <-c (mRNA=1, protein=1)
#Set the starting value for response variables
output <- as.data.frame(rk4(xstart, times, constitutiveexpression, parms))
plot(output$time, output$protein, ylim=c(0,2000), ylab="protein (molecules)",xlab="time (min)",type="n") }