User:Saroj Pandey/Notebook/SNP PCR optimization/2014/09/18: Difference between revisions
Saroj Pandey (talk | contribs) |
Saroj Pandey (talk | contribs) |
||
| Line 6: | Line 6: | ||
| colspan="2"| | | colspan="2"| | ||
<!-- ##### DO NOT edit above this line unless you know what you are doing. ##### --> | <!-- ##### DO NOT edit above this line unless you know what you are doing. ##### --> | ||
== | ==SNP PCR== | ||
'''Primers ending in G''' | |||
*fixed forward 161bp L: high 3' stability | |||
TAS2R38_1f: GGGATGTAGTGAAGAGGCAGG | |||
Length: 21 bp Tm: 61.0 °C GC: 57.1 % ANY: 2.0 SELF: 0.0 | |||
TAS2R38_1r: GATGGCTTGGTAGCTGTGGT | |||
Length: 20 bp Tm: 60.1 °C GC: 55.0 % ANY: 4.0 SELF: 0.0 | |||
Product Size: 161 bp Pair Any: 3.0 Pair End: 0.0 | |||
*fixed backward 235bp R: high end self complementarity | |||
TAS2R38_2f: GGTGGCAACCAGGTCTTTAG | |||
Length: 20 bp Tm: 59.6 °C GC: 55.0 % ANY: 6.0 SELF: 2.0 | |||
TAS2R38_2r: CAATCACTGTTGCTCAGTGC | |||
Length: 20 bp Tm: 58.0 °C GC: 50.0 % ANY: 6.0 SELF: 4.0 | |||
Product Size: 235 bp Pair Any: 5.0 Pair End: 1.0 | |||
'''Primers ending in C''' | |||
*fixed forward 161bp | |||
TAS2R38_C1f: GGGATGTAGTGAAGAGGCAGC | |||
Length:21 bp Tm: 61.2 °C GC:57.1 % ANY:3.0 SELF:3.0 | |||
TAS2R38_C1r: GATGGCTTGGTAGCTGTGGT | |||
Length:20 bp Tm: 60.1 °C GC:55.0 % ANY:4.0 SELF:0.0 | |||
2. | *fixed backward 235bp | ||
TAS2R38_C2f: GGTGGCAACCAGGTCTTTAG | |||
Length:20 bp Tm: 59.6 °C GC:55.0 % ANY:6.0 SELF:2.0 | |||
TAS2R38_C2f: CAATCACTGTTGCTCAGTGG | |||
Length:20 bp Tm: 57.8 °C GC:50.0 % ANY:6.0 SELF:4.0 | |||
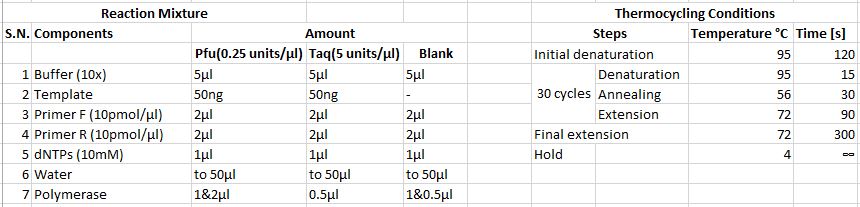
'''PCR 1''' | |||
[[Image:SNP_PCR1.JPG]] | |||
[[Image:Electrophoresis_GA_11.09.jpg|right|thumb|350px|Electrophoresis]] | |||
[[Image:Electrophoresis_GA_12.09.jpg|right|thumb|350px|Electrophoresis]] | |||
'''PCR 2''' | |||
[[Image:SNP_PCR2.JPG]] | |||
[[Image:Electrophoresis_GA_16.09.jpg|right|thumb|350px|Electrophoresis]] | |||
<!-- ##### DO NOT edit below this line unless you know what you are doing. ##### --> | |||
|} | |||
__NOTOC__ | |||
==PCR for gDNA amplification== | ==PCR for gDNA amplification== | ||
Revision as of 04:20, 19 September 2014
| <html><img src="/images/9/94/Report.png" border="0" /></html> Main project page <html><img src="/images/c/c3/Resultset_previous.png" border="0" /></html>Previous entry<html> </html>Next entry<html><img src="/images/5/5c/Resultset_next.png" border="0" /></html> | |
SNP PCRPrimers ending in G
TAS2R38_1f: GGGATGTAGTGAAGAGGCAGG Length: 21 bp Tm: 61.0 °C GC: 57.1 % ANY: 2.0 SELF: 0.0 TAS2R38_1r: GATGGCTTGGTAGCTGTGGT Length: 20 bp Tm: 60.1 °C GC: 55.0 % ANY: 4.0 SELF: 0.0 Product Size: 161 bp Pair Any: 3.0 Pair End: 0.0
TAS2R38_2f: GGTGGCAACCAGGTCTTTAG Length: 20 bp Tm: 59.6 °C GC: 55.0 % ANY: 6.0 SELF: 2.0 TAS2R38_2r: CAATCACTGTTGCTCAGTGC Length: 20 bp Tm: 58.0 °C GC: 50.0 % ANY: 6.0 SELF: 4.0 Product Size: 235 bp Pair Any: 5.0 Pair End: 1.0
Primers ending in C
TAS2R38_C1f: GGGATGTAGTGAAGAGGCAGC Length:21 bp Tm: 61.2 °C GC:57.1 % ANY:3.0 SELF:3.0 TAS2R38_C1r: GATGGCTTGGTAGCTGTGGT Length:20 bp Tm: 60.1 °C GC:55.0 % ANY:4.0 SELF:0.0
TAS2R38_C2f: GGTGGCAACCAGGTCTTTAG Length:20 bp Tm: 59.6 °C GC:55.0 % ANY:6.0 SELF:2.0 TAS2R38_C2f: CAATCACTGTTGCTCAGTGG Length:20 bp Tm: 57.8 °C GC:50.0 % ANY:6.0 SELF:4.0
PCR 1   PCR 2 
| |
PCR for gDNA amplification
Product Size: 992 bp Pair Any: 6.0 Pair End: 2.0
TAS2R38_gaF: ATCCGTGATGCTGTGCTATG
Length: 20 bp Tm: 59.7 °C GC: 50.0 % ANY: 4.0 SELF: 0.0
TAS2R38_gaR: GCATCCCAGAAGAAACCAGA
Length: 20 bp Tm: 60.2 °C GC: 50.0 % ANY: 2.0 SELF: 0.0
PCR 1
Agarose gel electrophoresis
- 0.7% agarose gel prepared
- 700 mg of agarose was added to water and heated at 600 °C for 90 seconds
- 10 mg gel red was added and mixed
- the gel mixture was added to the casting apparatus and comb was placed
- 450 ml TAE buffer was poured to the electrophoretic chamber
- After the gel was set, it was transferred to the electrophoretic chamber
- the comb was slowly taken out
- Sample mixtures (5μL water + 2μL loading dye + 5μL PCR product / 2μL DNA ladder) were loaded to individual wells and the chamber was covered
- electrophoresis was run at 120 V for 30 minutes

Observations
- first few samples including 100 bp ladder are invisible
- 1 kb ladder does not show distinct bands
- All the PCR products are seen below 250bp and are diffused
- Desired region of the genomic DNA was not amplified
PCR 2
- Amount of Phusion polymerase was changed
- Annealing temperature was decreased
- Taq DNA polymerase was also used


Agarose gel electrophoresis
- 0.7% agarose gel prepared
- 700 mg of agarose was added to water and heated at 600 °C for 90 seconds
- 10 mg gel red was added and mixed
- the gel mixture was added to the casting apparatus and comb was placed
- 450 ml TAE buffer was poured to the electrophoretic chamber
- After the gel was set, it was transferred to the electrophoretic chamber
- the comb was slowly taken out
- (5μL water + 2μL loading dye + 5μL PCR product / 2μL DNA ladder)
Sample mixtures were loaded to individual wells and the chamber was covered
- electrophoresis was run at 120 V for 30 minutes
Observation
- Phusion polymerase could not amplify the desired region of DNA
- Two bands were observed for Taq polymerase: one at about 1000bp and the other below 250bp
- Amplifiction was successful with taq DNA polymerase
- however, the bands are unclear and diffused
- reason for unclear and diffused bands may be the composition of agarose gel as it was prepared just in water
Agarose gel electrophoresis (repeated)
- agarose gel was prepared in TAE buffer and the samples were run
Observation
- bands look much clear and distinct
- PCR products of Taq DNA polymerase show bands at 1kbp which was estimated (992 bp)
- blank with Phusion DNA polymerase shows a thin band and that with Taq DNA polymerase shows a discinct and below 100 bp. These bands could not be primer dimers as they could only be as long as 40bp.
- unspecific bands/products were also seen in other samples
DNA concentration
1. T3: 520.9 ng/μL (260:280=1.79)
2. N3: 406.7 ng/μL (260:280=1.83)
- Samples T3 and N3 were sent for sequencing
|}