User:Vincent Rouilly: Difference between revisions
No edit summary |
|||
| Line 5: | Line 5: | ||
|style="background:#efefef" align="right"| | |style="background:#efefef" align="right"| | ||
<font size="4">Vincent Rouilly</font size> | <font size="4">'''Vincent Rouilly''', PhD Candidate</font size> | ||
<font size="2">Department of Bioengineering<br> | <font size="2">Department of Bioengineering<br> | ||
Imperial College London<br> | Imperial College London<br> | ||
South Kensington | South Kensington | ||
SW7 2AZ<br> | |||
London, UK<br> | London, UK<br> | ||
email: vincent. | email: vincent.rouilly03 (at) imperial.ac.uk | ||
</font size> | </font size> | ||
|} | |} | ||
== Education== | == Education and Past Work Experiences== | ||
*PhD candidate at Imperial College London | * PhD candidate at Imperial College London | ||
*M.Sc, ENST Paris and ERASMUS exchange at ETSIT Madrid. | * GE Medical Systems, Paris, Software R&D Engineer. | ||
* Visiospace, Paris, Software R&D Engineer. | |||
* M.Sc, ENST Paris and ERASMUS exchange at ETSIT Madrid (Signal and Image Processing Major). | |||
== Research Interests == | == Research Interests == | ||
* Computational Biology: stochastic processes, Petri Nets, Agent Based modelling. | * Computational Biology: stochastic processes, Generalized Petri Nets, Agent Based modelling, Design of Experiments. | ||
* Synthetic Biology and | * Contributing to build a Synthetic Biology framework (establishing standards, design for modularity and re-usability, parts characterization) + societal impact of Synthetic Biology. | ||
* Laboratory Information System. | * Laboratory Information System. | ||
* Lab Automation | |||
== Current Research Projects== | |||
* Writing-up PhD thesis: "An Engineering Approach To The Modeling Of Genetic Circuits" | |||
* iGEM instructor (2006, 2007) | |||
* Supervising master project: "Sensitivity analysis of genetic circuit induction protocols". | |||
* Supervising UROP project: "Building modular biobricks models implemented with CellML" | |||
== OWW Contributions == | == OWW Contributions == | ||
* Steering committee member and outreach co-chair. | |||
* [[Minicells | minicells]] | * [[Minicells | minicells]] | ||
* [[User:Vincent_Rouilly/Hands_On_BioBrick_Activity | Hands on BioBrick Activity for a science fair]] | * [[User:Vincent_Rouilly/Hands_On_BioBrick_Activity | Hands on BioBrick Activity for a science fair]] | ||
| Line 41: | Line 49: | ||
* [[IGEM:IMPERIAL/Methodology| iGEM@imperial project framework ]] | * [[IGEM:IMPERIAL/Methodology| iGEM@imperial project framework ]] | ||
* [[IGEM:IMPERIAL/2006| iGEM@imperial Instructor]] | * [[IGEM:IMPERIAL/2006| iGEM@imperial Instructor]] | ||
*[[IGEM:IMPERIAL/Methodology/Part_documentation_template | iGEM@imperial template to document a part]] | * [[IGEM:IMPERIAL/Methodology/Part_documentation_template | iGEM@imperial template to document a part]] | ||
*[[IGEM:IMPERIAL/Methodology/IGEM_WetLab_Planning_Tool | IGEM WetLab Planning Tool]] | * [[IGEM:IMPERIAL/Methodology/IGEM_WetLab_Planning_Tool | IGEM WetLab Planning Tool]] | ||
*[[OWW_Image_Editor | OWW Image Annotation Tool]] | * [[OWW_Image_Editor | OWW Image Annotation Tool]] | ||
Other: | Other: | ||
* [[IGEM:IMPERIAL/2006/Created_Pages]] | * [[IGEM:IMPERIAL/2006/Created_Pages]] | ||
== Interesting links found on OWW== | |||
*[[Endy:F2620 | F2620 Part Characterization (Labno/Canton)]] | |||
*[[Endy:Victor3_plate_reader/Calibrating_the_GFP-separated_label | GFP calibration and measurements]] | |||
== OWW ideas (random) == | == OWW ideas (random) == | ||
Revision as of 11:05, 14 July 2007
|
Vincent Rouilly, PhD Candidate Department of Bioengineering email: vincent.rouilly03 (at) imperial.ac.uk
|
Education and Past Work Experiences
- PhD candidate at Imperial College London
- GE Medical Systems, Paris, Software R&D Engineer.
- Visiospace, Paris, Software R&D Engineer.
- M.Sc, ENST Paris and ERASMUS exchange at ETSIT Madrid (Signal and Image Processing Major).
Research Interests
- Computational Biology: stochastic processes, Generalized Petri Nets, Agent Based modelling, Design of Experiments.
- Contributing to build a Synthetic Biology framework (establishing standards, design for modularity and re-usability, parts characterization) + societal impact of Synthetic Biology.
- Laboratory Information System.
- Lab Automation
Current Research Projects
- Writing-up PhD thesis: "An Engineering Approach To The Modeling Of Genetic Circuits"
- iGEM instructor (2006, 2007)
- Supervising master project: "Sensitivity analysis of genetic circuit induction protocols".
- Supervising UROP project: "Building modular biobricks models implemented with CellML"
OWW Contributions
- Steering committee member and outreach co-chair.
- minicells
- Hands on BioBrick Activity for a science fair
- Lab Automation Review
- Stochastic Gene Expression
- Registry of Standard Models
- Registry of Standard Models/Architecture
- Catalog of standard models in CellML
- Basic Components in CellML
- iGEM@imperial project framework
- iGEM@imperial Instructor
- iGEM@imperial template to document a part
- IGEM WetLab Planning Tool
- OWW Image Annotation Tool
Other:
Interesting links found on OWW
OWW ideas (random)
building a stronger community
- get people to define on their User page their area(s) of interest using 'wiki-categories' (could help to find collaborators).
- if people defines their area of interest using references, would be interesting to use that information to match people's profiles.
support to find relevant info
- 'contextual' browsing ('hacking' google-ads-type-of-service to show other relevant wiki-pages or wiki-users on the side of a wiki page ... instead of ads).
- set essential keywords pages. 'Experts', in a given field, would recommend a set of keywords to help new comers to google more efficiently.
- setting-up a calendar with info about conferences, workshops, submission deadlines ...
adding features
- implementing java applets which are enable to read/write an xml-like format into the wiki:
- image annotation (microscope images, gel analysis, colonies plate ..) see iNote, SVG.
- graph editor/viewer to build/view/run kinetic models (SBML, CellML ...)
Project
Here is an overview of the modelling framework I am working on:
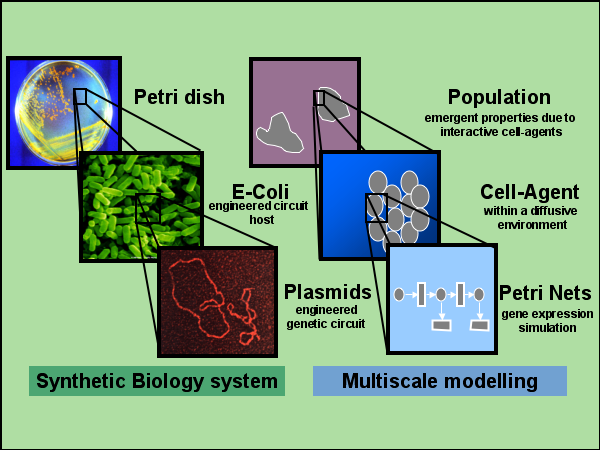
Who is visiting my page?
<html> <a href="http://clustrmaps.com/counter/maps.php?url=http://openwetware.org/wiki/User:Vincent" id="clustrMapsLink"><img src="http://clustrmaps.com/counter/index2.php?url=http://openwetware.org/wiki/User:Vincent" border=1 alt="Locations of visitors to this page"onError="this.onError=null; this.src='http://www.meetomatic.com/images/clustrmaps-back-soon.jpg'; document.getElementById('clustrMapsLink').href='http://clustrmaps.com/'"> </a>
<!-- Start of StatCounter Code --> <script type="text/javascript" language="javascript"> var sc_project=1775726; var sc_invisible=0; var sc_partition=16; var sc_security="e9fd94f4"; </script>
<script type="text/javascript" language="javascript" src="http://www.statcounter.com/counter/frames.js"></script><noscript><a href="http://www.statcounter.com/" target="_blank"><img src="http://c17.statcounter.com/counter.php?sc_project=1775726&java=0&security=e9fd94f4&invisible=0" alt="website tracking" border="0"></a> </noscript> <!-- End of StatCounter Code -->
</html>