User:Kara David/Notebook/Biology 210 at AU
3/19/15 Zebrafish Experiment
The purpose of the Zebrafish experiment was to study how different substances, in this case nicotine, affected the growth, development and behavior of Zebrafish throughout the embryonic and larval stages of their lives.
The materials required for this lab were forty Zebrafish embryos, twenty in a control group and twenty in an experimental group, water and liquid nicotine. The method used most for this lab was observation. However, the set-up did require some work. First, we got two different petri dishes and we filled the control dish up with 20 mLs of Deerpark water and the experimental dish was filled with 20 mLs of Deepark water and liquid nicotine. Then, we used a dropper to transfer 20 healthy translucent embryos into each dish. Around the fourth day, 10 mLs of water and any dead embryos were removed. Then, 25 mLs of fresh water was added and liquid nicotine was added to the experimental dish. The dead embryos were saved in paraformaldehyde. Around the seventh day,5 mLs of water and any dead embryos were removed and 5 mLs of fresh water along with nicotine (in the experimental dish only) was added. Some of the live embryos from each dish were preserved in paraformaldehyde. During the second week of our experiment, we continued to observe the Zebrafish and we removed 5 mLs of water and added 10 mLs of fresh water. At this time, we also used a dropper to feed the Zebrafish two drops of paramecium, which we also did every time we changed the water. At the end of the second week, we made our final observations and measurements and the surviving Zebrafish were collected and put in an aquarium for safekeeping.
Data and Observations
No pictures available.
The Zebrafish were observed multiple times throughout their development whilst being exposed to liquid nicotine. The first day we observed the fish, many of the embryos had died and they had to be replaced. The next time we observed them, many of them had developed into their larval stage but some were still embryos which could be due to the nicotine. The third time we observed the fish, mostly all of them had developed completely, some had died, and approximately two or three fish were still embryonic, which means they were probably dead. Also, some of the larval fish were dead and even some of the fully developed had died. However, the rest of the fully developed were alive and well - for the most part. Some of the fully developed Zebrafish seemed to not be affected by the nicotine. However, the other fully developed Zebrafish seemed to have been affected behaviorally by the nicotine, which was evident by their movement. Some moved very slowly and not very often and some only moved if they were prodded with a dropper. The last day we observed the fish, some more had died, there were only four left in the control group and two left in the experimental group in the end. However, the ones that did survive seemed to be the ones that were the strongest and fastest swimmers the last time we observed them.
It can be concluded that the nicotine had mostly negative effects on the Zebrafish and effects similar to those found in the papers that we read. Many of our Zebrafish died which could be from the nicotine exposure or from other causes but the slow growth and development and the slow and jittery movement is most likely due to the nicotine exposure.
3/05/15 PCR 16S Sequence
The purpose of this lab was to identify the type of bacteria that was found within our transect.
The materials required were our bacteria rRNA, the primers and polymerase used for our PCR reaction. The method used was PCR. To perform PCR, we selected one bacteria from our nutrient agar plate and one from our tetracycline plate. Each sample was labeled and then, we isolated the DNA from the bacteria in the colonies and used primers and PCR to selectively amplify the 16S rRNA gene. We started by transferring a single colony of bacteria to a tube containing 100 micro liters of water. The tube was then incubated at 100 degrees Celsius for 10 minutes in a heat block and then centrifuged at 13,400 rpm for 5 minutes. While our samples were being centrifuged, we added 20 micro liters of a primer and water mixture to a labeled PCR tube and mixed it together so the PCR bead could dissolve. Then, 5 micro liters of the supernatant from our centrifuged samples were added to the 16S PCR reaction and placed into the PCR machine. The next week in lab we ran the PCR products on an agarose gel to see if there was a PCR product. The PCR product was then purified by our instructors for sequencing. Once our samples were sequenced, we used a website to identify our bacteria.
Data and Observations
No picture available for PCR reaction.
Using GENEWIZ, a website that is used to sequence DNA, we found the raw sequences for our two samples. The raw sequences for our two samples, Sample A and Sample D can be seen below.
Sample A (MB65)
NNNNNNNNNNNNNNNNNCNNNNNNTGCNGNNNNANGGNNGNCNGNNNNNNANCAATCCTGGCGGCGAGTGGCGAACGGGT GAGTAATACATCGGAACGTGCCCAATCGTGGGGGATAACGCAGCGAAAGCTGTGCTAATACCGCATACGATCTACGGATG AAAGCAGGGGATCGCAAGACCTTGCGCGAATGGAGCGGCCGATGGCAGATTAGGTAGTTGGTGAGGTAAAGGCTCACCAA GCCTTCGATCTGTAGCTGGTCTGAGAGGACGACCAGCCACACTGGGACTGAGACACGGCCCAGACTCCTACGGGAGGCAG CAGTGGGGAATTTTGGACAATGGGCGAAAGCCTGATCCAGCCATGCCGCGTGCAGGATGAAGGCCTTCGGGTTGTAAACT GCTTTTGTACGGAACGAAACGGCCTTTTCTAATAAAGAGGGCTAATGACGGTACCGTAAGAATAAGCACCGGCTAACTAC GTGCCAGCAGCCGCGGTAATACGTAGGGTGCAAGCGTTAATCGGAATTACTGGGCGTAAAGCGTGCGCAGGCGGTTATGT AAGACAGTTGTGAAATCCCCGGGCTCAACCTGGGAACTGCATCTGTGACTGCATAGCTAGAGTACGGTAGAGGGGGATGG AATTCCGCGTGTAGCANTGNAATGCGTAGATATGCGGAGGAACACCGATGGCGAANGCAATCCCCTGGACCTGTACTGAC GCTCATGCACGAAAGCGTGGGGAGCAAACAGGATTAGATACCCTGGTAGTCCACGCCCTAAACGATGTCAACTGGTTGTT GGGTCTTCACTGACTCANTAACGAAGCTNACNCGTGAAGTTGACCGCCTGGGGAGTACGGCCGCAANGTTGAAACTCNAA NGAATTGACNNGGACCCGCACAAGCNGTGNATGATGTGNTTTAATTCNATGCAACGCGAAAACCTTACCCACCTTTGACA TGTACNNNANTTNNNCCAGANATGGCTTANTGCTCGAAANAAAANCGTAACNCANGTGCTNCATGNCTNNCGTCNNCNTC NTGTCGTGANA
Sample D (MB66)
NNNNNNNNNNNNNNNNNANANTGNANNCCNNAGCGGTAGCAGANGNTATCANGATGTCCGACAGCGGCTTGCNGATGAGG TACAAGTGTGGTTTATGCCTTTAGCCGGGGGAGGCACTTTCGTTGGGAAGATTACAACCCCATAATTATAATCGTGGCAT CTCTTGAAANGGACTGGTCCAGTGGAAAAAGAAGGGCCCGACCCTGATGANGCAGTTGGTACGGGGACGGTTCACCANGG CTGTGATGTTTGTGGGGCCTGANAGGGTGATCCCCCTGTGTGGTACGGAGACATTGACCCAACACCAATTGCAGGCGCCT CTGAGGAATATTGGACAATGGGTGAGAGCCTGATCNNNANTCNNCGNGAAGGATGACGGTGCTCCTGGTTGTATTCTTCT TTTGTATATTGATGGTGATTTCCTCGTGGGTGAAGCTGAATGAACTATACAAGCAGNAACCGGNGAGGCCCNTGCCTTCA GCCTCGGTNNTACNCAGGGTGTTGCCGTTTGAGAGATTTATTGNNTTNTCGAGGTTGGTTCNNGCNGANGGCNNACAATA TGCTGTANNNNTNACTNNNNGGTCAATCTGCATANGTTGGCGCGNGNCGCGACTNTTGGATATCTACCTTGCNTAAAANA NTCNNACANGGAANNCNTANATAATANCNNNNNCACCAATTGCGAANGCAGGTTACTATGTCTTAACTGACGCTGATGGA CGAAAGCGTGGGGAGCGAACAGGATTANATACCCTGGTANTCCACGCCNTNNNNNATGCTNACTCGTTTTTGGGNTCTTC NGATTCAGAGACTAAACNAAAGTGATAAGTTAGCCACCTGGGGAGTACGTTCNCAAGANTGAAACTCNAAGGAATTGACN GNNCCCGCACAANCGGNGGATTATGTGNNTTNATTCNATGATACGCNANGAANCCTTNNCCNANGCTTAANTGGGNANTN GATCGGTTTNNNANNNNACCTTNCCTTNNNCAATTTCAAGGTNCTGCATGGNTNGTCNNCNGCTNNNNCCNNNANTNNNA GNTAANTCCTGNNNNNNNGNNNCCCCNTGTCNCNNN
Then, using BLAST, we determined that our first sample was Variovorax, which is a gram-negative bacterium from the family Comamonadaceae. Variovorax has an important role in biodegradative processes in nature and can promote plant growth (Jamieson, Pehl, Gregory, Orwin, 2009). Our second sample was identified as Chryseobacterium, which is also a gram-negative bacterium and is from the family Flavobacteriaceae. Chryseobacterium is a species of bacteria that can typically be found in soil and water (Kirby, Sader, Walsh, Jones, 2004)
It can be concluded that both of our samples match our observed colony and morphology. Both of the strains of bacteria that we found are commonly found in soil which explains why we found them in our transect. Variovorax is a ubiquitous, gram-negative, aerobic bacterium typically found in soil that plays a very important role in common biodegradative processes that are seen throughout nature (Jamieson, Pehl, Gregory, Orwin, 2009). Chryseobacterium, also ubiquitous, gram-negative and typically found throughout nature, it is responsible for causing many diseases and is considered the most pathogenic in its genus (Kirby, Sader, Walsh, Jones, 2004). Chryseobacterium has also been found to be resistant to antibiotics like aminoglycosides, tetracyclines, chloramphenicol, erythromycin, clindamycin and teicoplanin (Kirby, Sader, Walsh, Jones, 2004) which was proven in our experiment because we found one on our tetracycline plate.
References
Jamieson, W., Pehl, M., Gregory, G., & Orwin, P. (2009). Coordinated Surface Activities In Variovorax Paradoxus EPS. BMC Microbiology, 9, 124-124.
Kirby, J., Sader, H., Walsh, T., & Jones, R. (2004). Antimicrobial Susceptibility and Epidemiology of a Worldwide Collection of Chryseobacterium spp.: Report from the SENTRY Antimicrobial Surveillance Program (1997-2001). Journal of Clinical Microbiology, 42, 445-448.
2/19/15 Vertebrates
The purpose of this was to observe the different types of vertebrates found in our transect and note the diversity of vertebrates found.
The material required was our textbook. The method used was observation of our transect and use of the textbook and internet to research the different vertebrates and ecological concepts noted in the lab manual.
Data and Observations
The different vertebrates observed in our transect were squirrels, chipmunks, rats, sparrows and robins. A food web was constructed based off of each of these vertebrates and based off the other biotic components found in our transect that these vertebrates would eat which can be seen below. Although they would not be used for food and therefore are not included in the food web, abiotic components in our transect can also benefit the vertebrates by acting as shelter like the lampposts, benches and trashcans which can be used to scavenge for food (mostly by the rats). However, there are also abiotic components found in our transect that can be harmful to the vertebrates like rat traps. Each vertebrate was classified by phylum, class, order, family and genus which can be found in Table 1: Vertebrate Classification below. The British Ecological Society defines community as all the species in a certain ecosystem (our transect) that interact with each other. These organisms represent community by feeding off the same biotic components found in the transect. The Sustainable Scale Project define carrying capacity as the maximum population of a species that can survive in a certain ecosystem (our transect). These organisms represent carrying capacity by there only being a certain amount of each species in our transect. Science Daily defines trophic levels essentially as what level in the food chain each organism is, what it eats and what eats it. The different vertebrates in our transect are mostly all at high trophic levels when in our transect because there are not many predators for large vertebrates found there.
Transect Three Vertebrate Food Web 
Table 1: Vertebrate Classification 
It can be concluded that the different vertebrates found in our transect are very diverse but also similar at the same time. Multiple vertebrates observed were mammalia and rodentia. The birds were both aves and passeriformes. They started to differ once it came down to which family and genus' they were classified into. However, two of the rodentia were classified as sciuridae. The different vertebrates were also similar in diets which can be seen in the food web above.
British Ecological Society. (2014, January 1). Retrieved February 19, 2015, from http://www.britishecologicalsociety.org/about-ecology/key-concepts-in-ecology/
Carrying Capacity. (2003, January 1). Retrieved February 19, 2015, from http://www.sustainablescale.org/ConceptualFramework/UnderstandingScale/MeasuringScale/CarryingCapacity.aspx
Trophic level. (2015, February 18). Retrieved February 19, 2015, from http://www.sciencedaily.com/articles/t/trophic_level.htm
2/19/15 Invertebrates
The purpose of this lab was to examine different types of invertebrates so we can understand the complexity or diversity of invertebrates and learn how the vertebrate organ systems evolved from the simplistic invertebrates.
The materials used for this lab were different invertebrate samples and microscopes. The methods used for this lab consisted mostly of observation. First, we had to observe a Planaria which is a type of a coelomate, then we observed nematodes which is a type of pseudocoelomate. Next, we observed an Annelida which is a type of coelomate. After this we observed different types of Arthropods from the five major classes: arachnida, diplopoda, chilopoda, insect and crustacea. Once we observed those, we broke down the Berlese Funnel and poured the top 10-15 mLs of liquid and organisms into one petri dish. The remaining liquid was poured into a second dish and then both were examined using a dissecting microscope. We then had to identify the class of Arthropoda found in our sample using an Insecta key and diagram of common soil invertebrates.
Data and Observations
The acoelomate, Planaria moved by contracting its body in and then stretching it out. It was flat and long. The pseudocoelomate, nematode was clear or colorless and small. It moved its body by wiggling and sliding forward. The coelomate, Annelida moved by squirming its body calmly using its entire body. It had no bones which caused its entire body to move at once. The invertebrates found in our sample were a springtail x, proturan x primitive, springtail x primitive, pseudoscorpion and soil mite. They ranged in size from 1 mm to 6 mm. The largest was the pseudoscorpion which measured at 6 mm and the smallest was the soil mite which measured at 1 mm. The organism that was most common was the springtail x primitive, it was found four times in the sample. Descriptions of the different invertebrates can be found in Table 1: Berlese Funnel Invertebrates and pictures (not ours) of what these organisms look like can be seen below. Drawings of what the organisms look like can also be found below in Figure 3: Common Soil Invertebrates.
Figure 3: Common Soil Invertebrates 
Table 1: Berlese Funnel Invertebrates 
Organism 2: Proturan X Primitive Insect 
Organism 3: Springtail X Primitive Insect 
Organism 4: Pseudoscorpion X LL 
It can be concluded that invertebrates are very diverse and that our transect includes a large variety of invertebrates.
2/12/15 Plantae and Fungi
The purpose of this lab was to examine the different types of plant life and fungi that can be found living in Transect Three so we can better understand the diversity of life found within our transect.
The materials used for this lab were plastic bags, a Berlese funnel, light fixture, tin foil, screen, flask and preservative inside the flask. The methods used for this lab were first collecting leaf litter, sticks and dirt and placing them in the first plastic bag to be used as a sample in lab. Then, samples from five different plants around our transect were collected and put into the second plastic bag to be used as a sample in lab. While collecting the samples pictures and notes for descriptive purposes were taken. Next was the setup of the Berlese Funnel which is when 25 mL of 50:50 ethanol/water solution was poured into the 50mL conical tube. The screening material was fit into the bottom of the funnel and taped so the leaf litter would not fall into the preservative. Then, the leaf litter was placed into the funnel. The funnel was then set up on a ring stand. Next, parafilm had to be placed around the funnel and conical tube so the ethanol would not evaporate which was also taped down. The funnel and conical tube were then placed approximately 1-2 inches underneath a 40 watt lamp and covered with foil and left to sit for the week.
Data and Observations
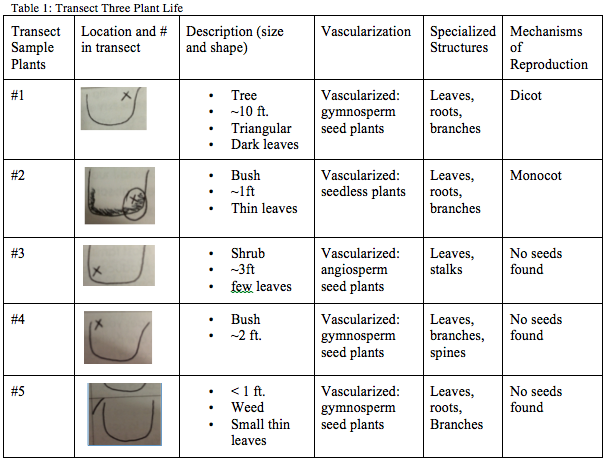
The leaf litter found in Transect Three was all primarily dead. Most of the leaves were medium or small sized leaves. It was all found on the ground surrounding the trees and bushes. Most of the leaf litter found was similar to each other in color, shape and size. There were only one or two bushes that had leaves still on them and those were bright green in color and there was little to no remaining leaves on the trees. Descriptions, locations, vascularization, specialization and mechanisms of the different plant samples can be found in Table 1: Transect Three Plant Life. The Fungi Sporangia are black, globe like structures with spores inside. They reproduce by releasing the spores when the sporangia open. There was no mold found in our transect or on our petri dishes from last week so we examined prepared slides with fungi on them. The first type of fungus we examined was Black Bread mold which can be classified as Zygomycota. The second type of fungus we examined was a cremini mushroom of Agaricus bisporus which can be classified as Basidiomucota.
Cremini Mushroom/Agaricus bisporus (Basidiomucota)

It can be concluded that Transect Three is not the most diverse transect. Considering all the life was somewhat similar. There was not a big diversity of life. It is possible that there will be a bigger diversity in later seasons considering most of the plants seem to be deciduous or it is possible that since most of the life in our transect was planted that there will never be a huge diversity in the life.
2/5/15 Microbiology and Identification of Bacteria with DNA Sequences
The purpose of this lab was to examine the different types of bacteria that were found in the sample taken from transect three using a microscope, wet mounts and gram stains.
The materials used for this lab were a microscope, different samples from the transect, dyes and decolorizers, iodine, safranin and intense heat (fire). The methods used for this lab were examining the different bacterium under the microscope, gram stain which is when the samples were dyed with different color dyes and decolorized to determine which bacteria were gram positive and which were gram negative, which is determined by a difference in cell wall.
Data and Observations
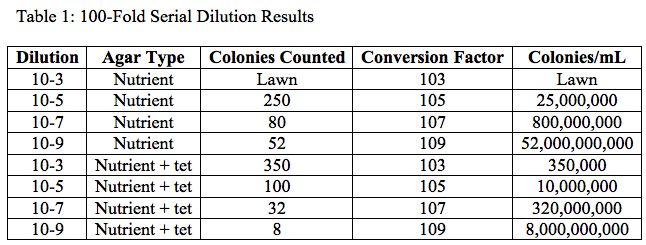
It can be concluded that the yellow and black/purple bacteria were affected by the tetracycline because there was only a little of the yellow bacteria found on the tet plates and there was none of the black/purple bacteria found.
1/29/15 Identifying Algae and Protists
The purpose of this lab was to learn how to identify different types of algae and protists and to learn how to use a dichotomous key to identify the different types, observe the Hay Infusion Cultures from last week and prepare serial dilutions for lab next week.
The materials used for this lab were a microscope, dichotomous key and different samples of different algae and protists. The methods used for this lab were viewing the organisms under the microscope and identifying them using a dichotomous key and taking two different samples from the Hay Infusion Cultures from the top of the jar and the bottom of the jar and making wet mounts which were then labeled. Then, the samples were both examined to see if there were any living organisms.
Data and Observations The algae identified was colpidium. It was oval in shape and measured to be 57 micrometers. The organisms identified in the transect sample were a Peranema which measured to be 20 micrometers and is a motile protozoa, an Amoeba proteus which measured to be 1200 micrometers and a Paramecium bursaria which measured to me 50 micrometers which were all found at the bottom of the sample. Only one organism could be found at the top of the transect sample and it was identified as a Spirostomum which is a type of protist and measured to be 2 mm.
It can be concluded that it was easier for life to grow at the bottom of the jar.
KD
1/21/15 Introduction to Biological Life at AU
The purpose of this lab was to learn more about the different biotic and abiotic components in different transects around the American University campus.
The materials used for this lab were a microscope, a jar used for storing the different abiotic and biotic components found at whichever transect each group was assigned to and water to mix with the components. The methods used for this lab were simply use of a microscope and collecting samples from the specific transect. After the sample was collected 10 grams of it was put into a jar along with 500 mL water and 0.1 gm of dried milk. The jar was then labeled and the contents had to be mixed together. After the sample, water and dried milk were mixed the jar was left to sit without the top on.
Data and Observations The abiotic components found in transect three were rat traps, snow, cement, lampposts and trashcans. The biotic components found in transect three were leaves, trees, plants, bushes, and different insects and animals like squirrels and birds.
It can be concluded that more life will grow from the sample taken at the transect and that over time the life at the actual transect will change due to climate changes. Once the weather gets warmer there will most likely be more biotic components seen at the transect.
KD
1/18/15 First Entry
This is my first entry on Open WetWare.
KD