IGEM:IMPERIAL/2009/Assays Protocols/Testing ConstructsOLD: Difference between revisions
James Field (talk | contribs) |
James Field (talk | contribs) |
||
| Line 47: | Line 47: | ||
<td><b>Link</b></td> | <td><b>Link</b></td> | ||
</tr> | </tr> | ||
<tr> | |||
<td></td> | |||
<td></td> | |||
<td></td> | |||
<td></td> | |||
<td></td> | |||
</tr> | |||
</table><br> | |||
<br> | |||
== Promoter Testing== | == Promoter Testing== | ||
Revision as of 07:49, 7 August 2009
Ligation Summary
- Running total of number of ligations required for the assembly of testing constructs.
1) Promoter Testing (4-5 ligations)
2) PAH (3 ligations)
3) Cellulase (3 ligations)
4) Thermoinduction (7 ligations)
5) Restriction Enzymes (12 ligations)
6)Encapsulation (15 ligations)
Running total = 45-46
1 LacIQ promoter BBa_K091111 Shipment: 00352 ID:941 Davidson 2 pSB1A2 A 100-300 LacIQ promoter
Testing Construct Summary
| Module | Part | BB # | Kit | Well # | Plasmid | Resistance | Copy # | Link |
Promoter Testing
We aim to use a generic promoter testing system, however we are unsure of the exact promoters that we are using at this stage becuase of the timer situation.
At this present moment in time, we are estimating using 4-5 different promoters in our system. Since each will need to be characterised, 4-5 ligations will be required to faciliate the assembly of the testing constructs.
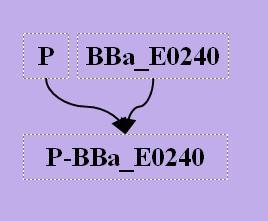
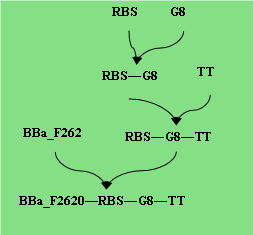
Testing Construct Assembly:
Input: POPs
Output: GFP
Parts Required: BBa_E0240, Each promoter to be characterised.
Module 1
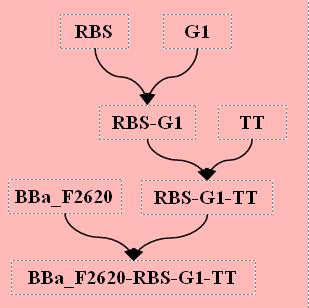
Testing Construct Assembly
We must test both cellulase and PAH, both testing constructs can be made using a total of 6 ligation steps.
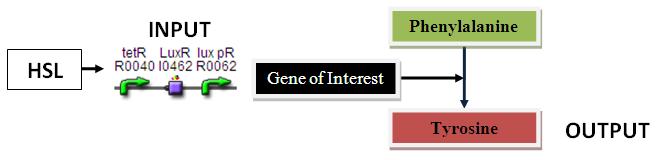
PAH Assay:
Input: HSL (POPs)
Output: PAH (Tyrosine)
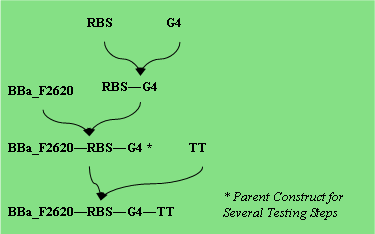
Parts Required: BBa_F2620, RBS, PAH, TT
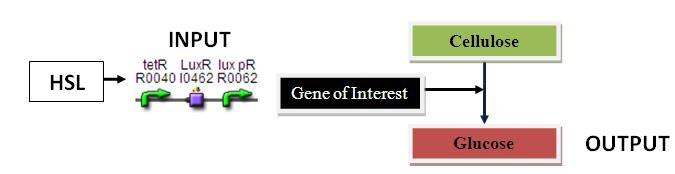
Cellulase Assay:
Input: HSL (POPs)
Output: Cellulase (Glucose)
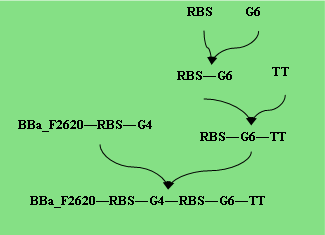
Parts Required: BBa_F2620, RBS, Cellulase, TT
Module 2
RcsB
- James Chappell 04:26, 4 August 2009 (EDT):I would join the RBS-Gx to the T first, think about the type of modules we want to submit to the registry. Think having a module that takes pops in and gives out G4 output is a good module to submit.
Inputs : HSL (POPs)
Outputs : Colanic Acid Production
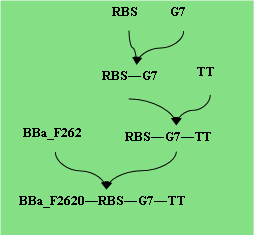
Parts: BBa_F2620, RBS, RcsB, TT
B30237
WaaL
Inputs : HSL (POPs)
Outputs : WaaL Ligase
Parts: BBa_F2620, RBS, WaaL, TT
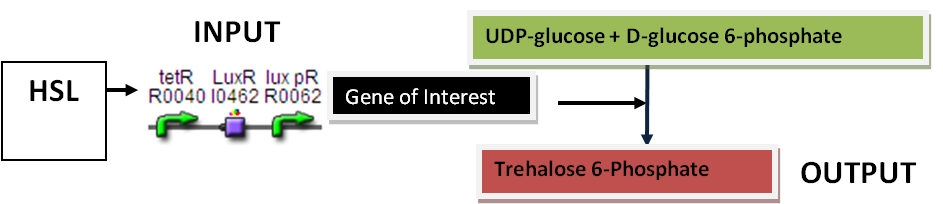
OtsA : Trehalose–6-Phosphate Synthase Assay
Inputs : HSL (POPs)
Outputs : Trehalose 6-phosphate Synthase
Parts: BBa_F2620, RBS, OtsA, TT
Assay Method
OtsB
Inputs : HSL (POPs)
Outputs : Trehalose 6-phosphate Phosphatase
Parts: BBa_F2620, RBS, OtsB, TT
Module 3
1)The thermoinduction system
Total number of ligations: 7
cI repressor
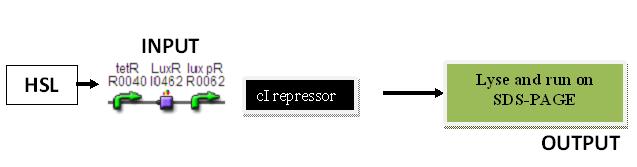
Input: HSL (F2620)
Output: Amount of cI protein
Plambda
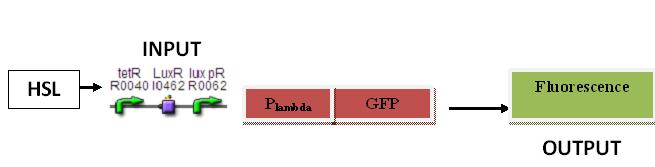
Input: HSL (F2620)
Output: Fluorescence
cI and Plambda

Input: HSL (F2620)
Output: (visual) cI physically binding to Plambda
cI and Plambda(GFP reporter)
Input: HSL (F2620)
Output: Change in fluorescence
2) Restriction Enzymes
CHANGE IMAGE - NO LONGER USING pBAD!
In total there will be 12 ligation steps.

We will test using a non-leaky pBAD promoter in order to utilise the GFP reporter cell death assay (see here).
An alternative to the GFP assay is to measure colony forming units.
DpnI
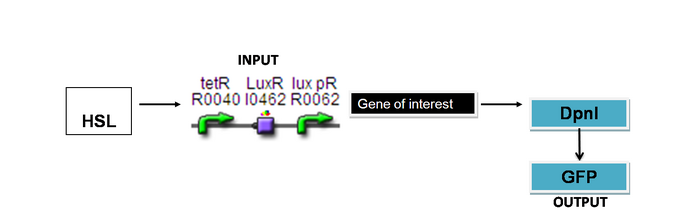
Input: HSL (F2620)
Output: Change in fluorescence
TaqI

Input: HSL (F2620)
Output: Change in fluorescence