BIO254:Transcription: Difference between revisions
mNo edit summary |
mNo edit summary |
||
| Line 39: | Line 39: | ||
CREB affects a variety of genes, including many immediate early genes (first set of genes to be turned on after a stimulus), such as c-Fos. CREB plays an interesting role in neuronal survival – while activation of CREB leads to neuronal survival, overactivation of CREB can lead to apoptosis. | CREB affects a variety of genes, including many immediate early genes (first set of genes to be turned on after a stimulus), such as c-Fos. CREB plays an interesting role in neuronal survival – while activation of CREB leads to neuronal survival, overactivation of CREB can lead to apoptosis. | ||
[[Image:ca2toCREB4.jpg|frame|Figure 1. Activation of CREB by Ca2+ Influx (Deisseroth 2003)]] | [[Image:ca2toCREB4.jpg|frame|Figure 1. Activation of CREB by Ca2+ Influx (Deisseroth 2003)]] | ||
<BR> | <BR> | ||
| Line 57: | Line 56: | ||
CBP: CREB binding protein<br> | CBP: CREB binding protein<br> | ||
CRE: cAMP response element<br> | CRE: cAMP response element<br> | ||
CaMKK: | CaMKK: CaM dependent protein kinase kinase <br> | ||
CaMKIV:<br> | CaMKII:CaM dependent protein kinase II<br> | ||
CaMKIV:CaM dependent protein kinase IV<br> | |||
NMDAR: N-methyl-D-aspartate receptor | NMDAR: N-methyl-D-aspartate receptor | ||
PKA: protein kinase A<br> | PKA: protein kinase A<br> | ||
| Line 67: | Line 67: | ||
GSK3: glycogen synthase kinase 3B<br> | GSK3: glycogen synthase kinase 3B<br> | ||
==History== | ==Research History of Activity-Dependent Transcription== | ||
In 1984, Stanton and Sarvey showed that long-term potentiation in the hippocampus can be blocked by inhibitors of protein synthesis. | In 1984, Stanton and Sarvey showed that long-term potentiation in the hippocampus can be blocked by inhibitors of protein synthesis. | ||
Revision as of 06:15, 24 October 2006
Introduction
In activity-dependent transcription, a postsynaptic neuron changes the level of transcription of specific genes in response to synaptic activity. While both long-term and short-term changes are produced by synaptic activity, long-term changes, such as synaptic plasticity and neuronal development, are often caused by changes in gene expression.
Activity-dependent transcription is critical for the proper function and development of the neural system including in LTP and memory formation, synaptic plasticity, neuronal survival, neurogenesis, dendritic arborization, and wiring.
General Mechanism for Activity-Dependent Transcription
Following are the steps leading to changes in gene expression:
A) Neuronal Activity
The presynaptic neuron excites or inhibits the postsynaptic neuron using neurotransmitters, such as glutamate or GABA.
B) Postsynaptic Response
The neurotransmitter activates metabotropic and/or ionotropic receptors activating G-protein/second messenger systems or producing ionic currents, respectively.
C) Signaling Pathways
Signaling proteins are activated (phosporylated/dephosphorylated) by either the G-protein pathways or the pathways affected by ionic concentrations (usually Ca2+ dependent). These pathways are often referred to as cascades because of the multiple proteins phosphorylated/dephosphorylated sequentially in a pathway.
D) Nuclear Localization & Transcription Factor Activation/Inactivation
An activated protein enters the nucleus and activates/inactivates an activating or repressing transcription factor. In some cases, the protein that enters the nucleus is itself an activated transcription factor which goes directly and binds to its specific DNA sequence.
E) Gene expression Increased/Decreased
Depending on whether the transcription factor is activated or inactivated and whether the transcription factor itself activates or represses gene expression, the transcription of the target gene(s) will be changed.
Transcription Factors
Along with protein-coding regions, the gene's DNA includes sequences that are involved in the control of the gene's expression. These sequences are called regulatory elements; transcription factors are usually sequence specific and bind to regulatory elements containing its binding sequence (~7-12 base pairs long). For example, CREB binds to the DNA sequence CRE (cAMP response element). In eukaryotes, the transcription of DNA to mRNA is carried out by RNA Polymerase II, which binds to the gene's promoter region. Transcription factors affect the ability of the polymerase to bind to the promoter region, thereby influencing the level of expression of the gene. Often, adaptor proteins (which interact with both the transcription factor and the polymerase) are required for the transcription factor (bound to its binding sequence within several hundred base pairs from the promoter region) to influence gene expression.
Examples of Pathways that Affect Transcription
Activation of CREB by Ca2+ Influx
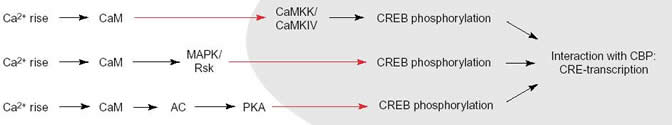
CREB, the cyclic AMP response element binding protein, is a transcription factor involved in various signaling pathways influencing synaptic plasticity, neuronal survival, and neuronal development. CREB remains constitutively bound to its DNA binding sequence, CRE; CREB must be phosphorylated on Ser-133 and bound to its activator CBP to become active and increase or decrease the expression of specific genes. The pathway that leads to CREB activation initially requires an increase in Ca2+ concentration, the activation of G-protein coupled receptors, or the binding of neurotrophins to cell receptors. For Ca2+ to enter the postsynaptic neuron, an excited presynaptic neuron has to release glutamate, which binds glutamate receptors (AMPA, NMDA, Kainate) and depolarize the postsynaptic neuron. In hippocampal neurons (and possibly in other neurons), Ca2+ ion flow through the NMDA receptor and the L-type voltage-gated calcium channel are required for the signaling cascade to begin. In other words, Ca2+ increases by themselves are not sufficient to activate CREB (Deisseroth 2003). This could be because L-type Ca2+ channel seems to carry a CaM binding site and the NMDA receptor has a calcium microdomain near it.
Four Ca2+ ions then bind to calmodulin forming a Ca2+/calmodulin complex. The complex activates CaMKII outside the nucleus and also translocates into the nucleus to activate CaM dependent protein kinase kinase (CaMKK). This protein then activates CamKIV which phosphorylates CREB at Ser-133. An adaptor protein, CBP then binds to CREB and the RNA polymerase.
CREB affects a variety of genes, including many immediate early genes (first set of genes to be turned on after a stimulus), such as c-Fos. CREB plays an interesting role in neuronal survival – while activation of CREB leads to neuronal survival, overactivation of CREB can lead to apoptosis.
More Images of CREB Pathway
UT Houston Simple picture of neurotransmitter activation of G-protein, increase of cAMP concentration, activation of PKA, and activation of CREB
Humboldt University, Berlin Simple picture of pathway from Ca2+ increase to CREB activation
Panomics Simple picture of various pathways influencing CREB
Eurekah Small picture that shows effect of neurotransmitter, Ca2+ increase, and growth factors on CREB
Sigma Aldrich CREB activation by growth factors
Biocarta Complicated picture of various pathways influencing CREB
Abbreviations
CaM: calmodulin
Ca2+/CaM: calcium calmodulin (calcium bound to calmodulin)
CaMK: calmodulin-dependent protein kinase
CREB: Ca2+/cAMP responsive element binding protein
CBP: CREB binding protein
CRE: cAMP response element
CaMKK: CaM dependent protein kinase kinase
CaMKII:CaM dependent protein kinase II
CaMKIV:CaM dependent protein kinase IV
NMDAR: N-methyl-D-aspartate receptor
PKA: protein kinase A
ERK: extracellular regulated kinase
MEK: MAPK kinase
ROS: reactive oxygen species
PI3-K: phopshatidyl inositol3 kinase
GSK3: glycogen synthase kinase 3B
Research History of Activity-Dependent Transcription
In 1984, Stanton and Sarvey showed that long-term potentiation in the hippocampus can be blocked by inhibitors of protein synthesis.
In 1986, Morgan and Curran studied the relationship between the Ca2+ influx and gene expression, specifically of c-fos. They suggested that Ca2+ would enter through the L-type Ca2+ channel and bind to calmodulin. Then, a calmodulin sensitive kinase would phosphorylate a transcription factor, which would then move into the nucleus.
In 1988, Sheng et. al. found a regulatory sequence upstream of c-Fos, which would turn out to be the CRE where CREB binds.
In 1989, Gonzalez and Montminy discovered that CREB needed to be phosphorylated at serine 133 to become active.
In 2000, Eric Kandel received a Nobel Prize in Physiology or Medicine partly for his work showing the importance of activity-dependent transcription in memory formation.
Notes
This article has focused on activity-dependent transcription in neurons. Activity-dependent transcription occurs in various excitable cells, such as muscle cells (ie, Wamhoff 2006).
Also, it is important to note that phosphorylation of a protein activates some proteins while it inactivates others.
References
Curtis J, Finkbeiner S. Sending signals from the synapse to the nucleus: possible roles for CaMK, Ras/ERK, and SAPK pathways in the regulation of synaptic plasticity and neuronal growth. J Neurosci Res. 1999 Oct 1;58(1):88-95.
Deisseroth K, Heist EK, Tsien RW. Translocation of calmodulin to the nucleus supports CREB phosphorylation in hippocampal neurons. Nature. 1998 Mar 12;392(6672):198-202.
Deisseroth K, Mermelstein PG, Xia H, Tsien RW. Signaling from synapse to nucleus: the logic behind the mechanisms.Curr Opin Neurobiol. 2003 Jun;13(3):354-65.
Gonzalez GA, Montminy MR. AMP stimulates somatostatin gene transcription by phosphorylation of CREB at serine 133. Cell. 1989 Nov 17;59(4):675-80.
Hardingham GE, Arnold FJ, Bading H. A calcium microdomain near NMDA receptors: on switch for ERK-dependent synapse-to-nucleus communication. Nat Neurosci. 2001 Jun;4(6):565-6.
Hardingham GE, Cruzalegui FH, Chawla S, Bading H. Mechanisms controlling gene expression by nuclear calcium signals. Cell Calcium. 1998 Feb-Mar;23(2-3):131-4.
Johannessen M, Delghandi MP, Moens U. What turns CREB on? Cell Signal. 2004 Nov;16(11):1211-27.
Morgan JI, Curran T. Role of ion flux in the control of c-fos expression. Nature 1986, 322:552-555.
Sheng M, Dougan ST, McFadden G, Greenberg ME. Calcium and growth factor pathways of c-fos transcriptional activation require distinct upstream regulatory sequences. Mol Cell Biol. 1988 Jul;8(7):2787-96.
Silva AJ, Kogan JH, Frankland PW, Kida S. CREB and memory. Annu Rev Neurosci. 1998;21:127-48.
Stanton PK, Sarvey JM. Blockade of long-term potentiation in rat hippocampal CA1 region by inhibitors of protein synthesis. J Neurosci. 1984 Dec;4(12):3080-8.
Squire LR, et al. Fundamental Neuroscience. Academic Press, Boston, 2003.
Wamhoff BR, Bowles DK, Owens GK. Excitation-transcription coupling in arterial smooth muscle. Circ Res. 2006 Apr 14;98(7):868-78.
Recent updates to the site:
List of abbreviations:
- N
- This edit created a new page (also see list of new pages)
- m
- This is a minor edit
- b
- This edit was performed by a bot
- (±123)
- The page size changed by this number of bytes
2 April 2026
|
|
19:57 | (Upload log) [Xinyu Liu; David Altman (8×)] | |||
|
|
19:57 David Altman talk contribs uploaded File:LabPicSpring2026.jpeg | ||||
|
|
19:57 David Altman talk contribs uploaded File:OASposterSpring2026.jpeg | ||||
|
|
19:56 David Altman talk contribs uploaded File:OAStalkSpring2026.jpeg | ||||
|
|
19:42 David Altman talk contribs uploaded File:Soto WUbites Spring 2026.pdf | ||||
|
|
19:42 David Altman talk contribs uploaded File:Shipp WUbites Spring 2026.pdf | ||||
|
|
19:42 David Altman talk contribs uploaded File:Margerum WUbites Spring 2026.pdf | ||||
|
|
19:42 David Altman talk contribs uploaded File:Bentley WUbites Spring 2026.pdf | ||||
|
|
19:40 David Altman talk contribs uploaded File:Bashioum WUbites Spring 2026.pdf | ||||
|
|
09:27 Xinyu Liu talk contribs uploaded File:Small2026.jpg | ||||
| 19:56 | Altman:Pictures diffhist +354 David Altman talk contribs | ||||
| 19:50 | Altman diffhist −54 David Altman talk contribs (→News) | ||||
|
|
19:40 | Altman:WUbites 2 changes history +992 [David Altman (2×)] | |||
|
|
19:40 (cur | prev) +40 David Altman talk contribs (→WUbites) | ||||
|
|
19:39 (cur | prev) +952 David Altman talk contribs (→WUbites) | ||||
| 18:36 | CHIP:Talks diffhist −19 Gabor Balazsi talk contribs | ||||
|
|
09:28 | Gao lab:Publications 2 changes history +549 [Xinyu Liu (2×)] | |||
|
|
09:28 (cur | prev) +24 Xinyu Liu talk contribs (→Publications) | ||||
|
|
09:26 (cur | prev) +525 Xinyu Liu talk contribs (→Publications) | ||||
1 April 2026
|
|
11:32 | Hu 2 changes history +121 [Hugangqing (2×)] | |||
|
|
11:32 (cur | prev) +2 Hugangqing talk contribs | ||||
|
|
11:32 (cur | prev) +119 Hugangqing talk contribs | ||||