Hugheslab: Difference between revisions
From OpenWetWare
Jump to navigationJump to search
No edit summary |
No edit summary |
||
| Line 3: | Line 3: | ||
<br> | <br> | ||
<p style="width:750px;"> | <p style="width:750px;"> | ||
<font size=" | <font size="4"> | ||
Our | Our laboratory studies circadian rhythms as a model to better understand how the nervous system regulates | ||
behavior and physiology. We are especially interested in uncovering the molecular mechanisms | behavior and physiology. We are especially interested in uncovering the molecular mechanisms | ||
driving rhythms of gene expression | driving rhythms of gene expression and determining how these molecular rhythms ultimately influence | ||
physiological rhythms. We use a combination of approaches, including behavioral neuroscience, genetics, | physiological rhythms. We use a combination of approaches, including behavioral neuroscience, genetics, | ||
genomics, and bioinformatics. | genomics, and bioinformatics. | ||
Revision as of 02:37, 8 January 2013
Home Lab Members Research JTK_Cycle MuscleDB Publications Protocols Academic Analytics Open Positions Contact
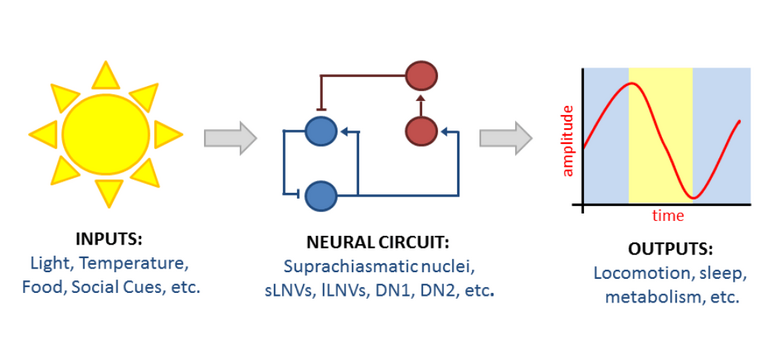
Our laboratory studies circadian rhythms as a model to better understand how the nervous system regulates behavior and physiology. We are especially interested in uncovering the molecular mechanisms driving rhythms of gene expression and determining how these molecular rhythms ultimately influence physiological rhythms. We use a combination of approaches, including behavioral neuroscience, genetics, genomics, and bioinformatics.

Department of Biology
University of Missouri, St. Louis
(Starting in August, 2013)