Nicolette S. Harmon Week 14: Difference between revisions
From OpenWetWare
Jump to navigationJump to search
| Line 33: | Line 33: | ||
#regulation of cellular biosynthetic process: Any process that modulates the frequency, rate or extent of the chemical reactions and pathways resulting in the formation of substances, carried out by individual cells [http://amigo.geneontology.org/cgi-bin/amigo/term_details?term=GO:0031326&session_id=7064amigo1323124805 The Gene Ontology] | #regulation of cellular biosynthetic process: Any process that modulates the frequency, rate or extent of the chemical reactions and pathways resulting in the formation of substances, carried out by individual cells [http://amigo.geneontology.org/cgi-bin/amigo/term_details?term=GO:0031326&session_id=7064amigo1323124805 The Gene Ontology] | ||
#regulation of biosynthetic process: Any process that modulates the frequency, rate or extent of the chemical reactions and pathways resulting in the formation of substances [http://amigo.geneontology.org/cgi-bin/amigo/term_details?term=GO:0009889&session_id=3245amigo1323124873 The Gene Ontology] | #regulation of biosynthetic process: Any process that modulates the frequency, rate or extent of the chemical reactions and pathways resulting in the formation of substances [http://amigo.geneontology.org/cgi-bin/amigo/term_details?term=GO:0009889&session_id=3245amigo1323124873 The Gene Ontology] | ||
#regulation of macromolecule metabolic process: Any process that modulates the frequency, rate or extent of the chemical reactions and pathways involving macromolecules | #regulation of macromolecule metabolic process: Any process that modulates the frequency, rate or extent of the chemical reactions and pathways involving macromolecules. [http://amigo.geneontology.org/cgi-bin/amigo/term_details?term=GO:0060255&session_id=6216amigo1323124944 The Gene Ontology] | ||
[[Image:Decreasedmicroarrays.gif]] | [[Image:Decreasedmicroarrays.gif]] | ||
Revision as of 02:27, 6 December 2011
DNA Microarray Project
Methods
Files
Microarray Statistics Text File
MAPPFinder Results(Increased) Text File
MAPPFinder Results(Increased) SpreadSheet
MAPPFinder Results(Decreased) Text File
MAPPFinder Results(Decreased) SpreadSheet
Table of Significant Genes for 2.5-50 Saturation
p< Number of Genes 0.05 215 0.01 42 0.001 3
GO Terms
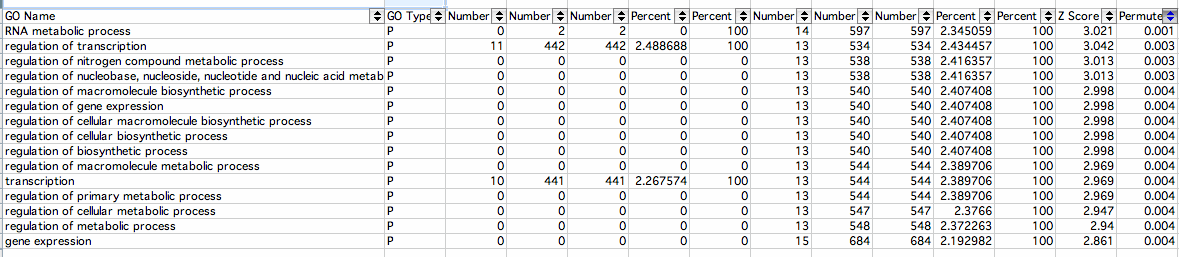
Increased:
- RNA metabolic process: The cellular chemical reactions and pathways involving RNA, ribonucleic acid, one of the two main type of nucleic acid, consisting of a long, unbranched macromolecule formed from ribonucleotides joined in 3',5'-phosphodiester linkage The Gene Ontology
- regulation of transcription: Any process that modulates the frequency, rate or extent of cellular DNA-dependent transcription The Gene Ontology
- regulation of nitrogen compound metabolic process: Any process that modulates the frequency, rate or extent of the chemical reactions and pathways involving nitrogen or nitrogenous compounds The Gene Ontology
- regulation of nucleobase, nucleoside, nucleotide and nucleic acid metabolic process: Any cellular process that modulates the frequency, rate or extent of the chemical reactions and pathways involving nucleobases, nucleosides, nucleotides and nucleic acids The Gene Ontology
- regulation of macromolecule biosynthetic process: Any process that modulates the rate, frequency or extent of the chemical reactions and pathways resulting in the formation of a macromolecule. The Gene Ontology
- regulation of gene expression: Any process that modulates the frequency, rate or extent of gene expression. This includes the production of an RNA transcript as well as any processing to produce a mature RNA product or an mRNA (for protein-coding genes) and the translation of that mRNA into protein The Gene Ontology
- regulation of cellular macromolecule biosynthetic process: Any process that modulates the frequency, rate or extent of cellular macromolecule biosynthetic process The Gene Ontology
- regulation of cellular biosynthetic process: Any process that modulates the frequency, rate or extent of the chemical reactions and pathways resulting in the formation of substances, carried out by individual cells The Gene Ontology
- regulation of biosynthetic process: Any process that modulates the frequency, rate or extent of the chemical reactions and pathways resulting in the formation of substances The Gene Ontology
- regulation of macromolecule metabolic process: Any process that modulates the frequency, rate or extent of the chemical reactions and pathways involving macromolecules. The Gene Ontology
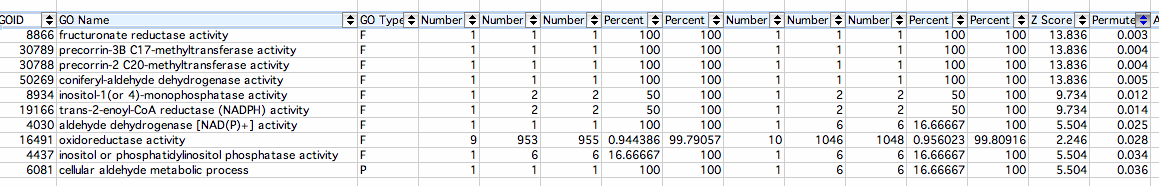
Decreased:
- fructuronate reductase activity: Catalysis of the reaction: D-mannonate + NAD(+) = D-fructuronate + H(+) + NADH The Gene Ontology
- precorrin-3B C17-methyltransferase activity: Catalysis of the reaction: S-adenosyl-L-methionine + precorrin-3B = S-adenosyl-L-homocysteine + precorrin The Gene Ontology
- precorrin-2 C20-methyltransferase activity:Catalysis of the reaction: S-adenosyl-L-methionine + precorrin-2 = S-adenosyl-L-homocysteine + H(+) + precorrin-3A The Gene Ontology
- coniferyl-aldehyde dehydrogenase activity: Catalysis of the reaction: coniferyl aldehyde + H2O + NAD(P)+ = ferulate + NAD(P)H + H+ The Gene Ontology
- inositol-1(or 4)-monophosphatase activity: Catalysis of the reaction: myo-inositol phosphate + H2O = myo-inositol + phosphate The Gene Ontology
- trans-2-enoyl-CoA reductase (NADPH) activity: Catalysis of the reaction: acyl-CoA + NADP+ = trans-2,3-dehydroacyl-CoA + NADPH + H+ The Gene Ontology
- aldehyde dehydrogenase [NAD(P)+] activity: Catalysis of the reaction: an aldehyde + NAD(P)+ + H2O = an acid + NAD(P)H + H+ The Gene Ontology
- oxidoreductase activity:The catalysis of redox reactions specifically relating to energy functions such as NAD+/NADH The Gene Ontology
- inositol or phosphatidylinositol phosphatase activity: Catalysis of the reaction: inositol phosphate(n) + H2O = inositol phosphate(n-1) + phosphate. This reaction is the removal of a phosphate group from an inositol phosphate The Gene Ontology
- cellular aldehyde metabolic process: The chemical reactions and pathways involving aldehydes, any organic compound with the formula R-CH=O, as carried out by individual cells The Gene Ontology
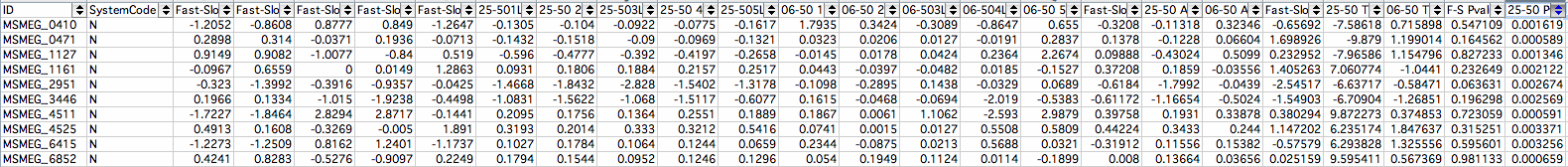
MSMEG IDs
- 5MSMEG_0410: MmpL Protein; P-Value: 0.001619 membrane protein involved in lipid transport
- 1MSMEG_0471: Transcriptional Regulator LysR Family Protein; P-Value: 0.000589
- 4MSMEG_1127: Probable Conserved Transmembrane Protein; P-Value: 0.001346
- 6MSMEG_1161: Taurine Transport System Permease Protein TauC; P-Value: 0.002122
- 8MSMEG_2951: [2Fe-2S] Binding Domain Protein; P-Value: 0.002674 Ferredoxin reducing/oxidizing enzyme powers TCA cycle independent of NAD/NADH
- 7MSMEG_3446: Hypothetical Protein; P-Value: 0.002569
- 2MSMEG_4511: Linear Gramicidin Synthetase mbtf; P-Value: 0.000591 antibiotic cells produces to kill other cells
- 10MSMEG_4525: Putative Oxygen-Independent Coproporhyrinogen III Oxidase; P-Value: 0.003371
- 9MSMEG_6415: Conserved Hypothetical Protein; P-Value: 0.003256
- 3MSMEG_6852: Putative Carboxylesterase/Lipase; P-Value: 0.000659 breaking down lipids
Links
BIOL368/F11:Class Journal Week 1
BIOL368/F11:Class Journal Week 2
BIOL368/F11:Class Journal Week 3
BIOL368/F11:Class Journal Week 4
BIOL368/F11:Class Journal Week 5
BIOL368/F11:Class Journal Week 6
BIOL368/F11:Class Journal Week 7
BIOL368/F11:Class Journal Week 8
BIOL368/F11:Class Journal Week 9
BIOL368/F11:Class Journal Week 10
BIOL368/F11:Class Journal Week 11
BIOL368/F11:Class Journal Week 12