IGEM:IMPERIAL/2006/project/Oscillator/project browser/Test Sensing Predator Construct/Implementation: Difference between revisions
From OpenWetWare
Jump to navigationJump to search
No edit summary |
|||
| Line 54: | Line 54: | ||
==Quality control== | ==Quality control== | ||
* | *These parts were sequenced to determine the correct sequence | ||
__NOTOC__ | __NOTOC__ | ||
| Line 63: | Line 61: | ||
!height="25pt" width="80pt"| Part !! Built !! Sequencing Results !! Primers Used for Sequencing | !height="25pt" width="80pt"| Part !! Built !! Sequencing Results !! Primers Used for Sequencing | ||
|- | |- | ||
|Put part logo and name here || | |Put part logo and name here || Yes || sequencing analysis and link to sequence results || PSBA2 primers | ||
|- | |- | ||
|Put part logo and name here || | |Put part logo and name here || Yes || sequencing analysis and link to sequence results || PSBA2 primers | ||
|- | |- | ||
|} | |} | ||
Revision as of 08:36, 28 October 2006
| Super Parts | Predator Construct | |||
|---|---|---|---|---|
| Actual Part | ||||
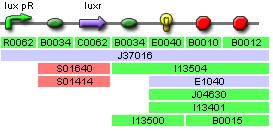
| Sub Parts | <bbpart>R0062</bbpart> | <bbpart>B0034</bbpart> | <bbpart>C0062</bbpart> | <bbpart>I13504</bbpart> |
Predator Test Construct
These two parts are very similar and have the same inputs and outputs, therefore they can be analysed in exactly the same way
This construct is used to characterise the pLuxR transfer function, and noteably its effect on YFP expression, which would be equivalent to AiiA expression in the full system with one promoter.
This construct is used to characterise the pLuxR transfer function, and noteably its effect on YFP expression, which would be equivalent to AiiA expression in the full system with two promoters.
The input is HSL and the outputs are LuxR and GFP. GFP will be mesured
Quality control
- These parts were sequenced to determine the correct sequence
| Part | Built | Sequencing Results | Primers Used for Sequencing |
|---|---|---|---|
| Put part logo and name here | Yes | sequencing analysis and link to sequence results | PSBA2 primers |
| Put part logo and name here | Yes | sequencing analysis and link to sequence results | PSBA2 primers |