User:Steven J. Koch/Notebook/Kochlab/2009/09/15/EarI pBR322: Difference between revisions
From OpenWetWare
Jump to navigationJump to search
(New page: ~~~~: Using restriction cross-correlate software & pBR322 sequence from genbank (http://www.neb.com/nebecomm/tech_reference/restriction_enzymes/sequences/GenBank/pBR322.gbk.txt), I find tw...) |
No edit summary |
||
| Line 1: | Line 1: | ||
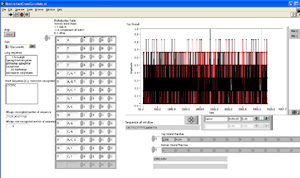
[[User:Steven J. Koch|Steve Koch]] 20:29, 15 September 2009 (EDT): Using restriction cross-correlate software & pBR322 sequence from genbank (http://www.neb.com/nebecomm/tech_reference/restriction_enzymes/sequences/GenBank/pBR322.gbk.txt), I find two [http://www.neb.com/nebecomm/products/productR0528.asp EarI] sites: | [[User:Steven J. Koch|Steve Koch]] 20:29, 15 September 2009 (EDT): Using restriction cross-correlate software & pBR322 sequence from genbank (http://www.neb.com/nebecomm/tech_reference/restriction_enzymes/sequences/GenBank/pBR322.gbk.txt), I find two [http://www.neb.com/nebecomm/products/productR0528.asp EarI] sites: | ||
[[Image:PBR322, EarI sites.png|right|thumb|EarI sites on pBR322]] | |||
* 2350 | * 2350 | ||
* 4154 | * 4154 | ||
Revision as of 17:31, 15 September 2009
Steve Koch 20:29, 15 September 2009 (EDT): Using restriction cross-correlate software & pBR322 sequence from genbank (http://www.neb.com/nebecomm/tech_reference/restriction_enzymes/sequences/GenBank/pBR322.gbk.txt), I find two EarI sites:
- 2350
- 4154
Thus, should produce two fragments, one approx 2556; the other approx 1804 bp. From the top strand, the overhangs read:
- 2350
- GCT (This is the overhang we have on our adapter duplex)
- 4154
- TTT
If we gel extract the big piece (approx 2556), it will be complementary to a 5' GCT overhang. So, I think we are all good. This is consistent I think with my earlier work (see below).
{{#widget:Google Documents |key=0ARLNnjMk2r_qZGdxamtoNnBfMTE4ZHBqOXFqZnY |width=500 |height=300 }}