BioMicroCenter:SPRI-Works
| HOME -- | SEQUENCING -- | LIBRARY PREP -- | HIGH-THROUGHPUT -- | COMPUTING -- | DATA MANAGEMENT -- | OTHER TECHNOLOGY |
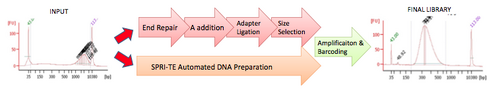
Fragmented samples submitted to the BioMicro Center are processed using the Beckman Coulter SPRIworks. This system preforms the sample cleanup, a overhang, ligation and preliminary size selection needed for creating Illumina libraries. The system accepts all amounts of insert and has been extensively tested.
The SPRI-te Nucleic Acid Extractor is a fully-automated library preparation system for Illumina sequencing and supports all Illumina applications except for smallRNA-Seq. It performs all of the traditional Illumina protocol steps listed on the Illumina Library Preparation page with the exception of PCR enrichment. A total of 10 libraries may be prepared simultaneously in 5 hours.
SPRIworks
|
The SPRI-Works system consists of three main components: the SPRI-TE Nucleic Acid Extractor, a method card that contains the liquid handling program, and a cartridge containing all of the reagents required to prepare a single library. The platform utilizes AMPureXP DNA-binding magnetic beads to purify fragmented DNA and perform size selection, eliminating the need for column purification and gel-based size selection. After the adapter ligation and size selection steps are complete, samples are ready for PCR enrichment and clean-up to complete the library prep process. For more information on the SPRI-Works system please visit www.spriworks.com. |
 |
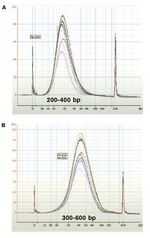
Size Selection
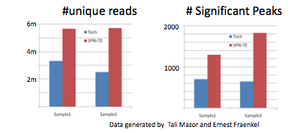
Library Complexity
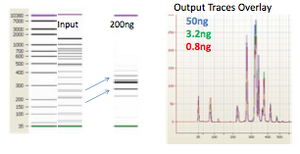
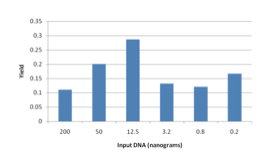
Effect of Concentration
Sample Submission
We suggest that you submit as much volume as possible since the first step of the protocol brings all sample volumes up to 100uL in water. For optimal yield, libraries should be fragmented so that the average fragment size is approximately 60nt shorter than the chosen size range to allow for the addition of partial adapters.
All samples submitted for BMC Fragmented DNA sample prep (SPRI-only) must be submitted with user supplied adapter mix. Adapter mixes should be pre-diluted and should have enough volume for 10uL of adapter per sample. For recommended dilutions for various sample concentrations, please see the chart below:
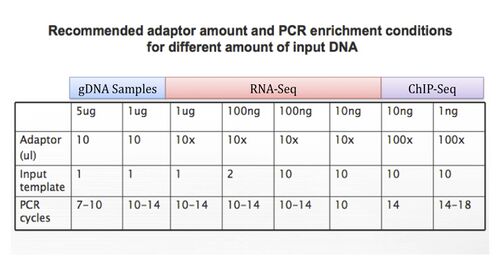
Please note that SPRI-only samples will need to be re-submitted separately if you wish to sequence with us later on.