Biomod/2013/Aarhus/Achievements And Future Work
<html> <style> /* ul.menu li.</html>Achievements And Future Work<html> a {
color: cyan;
}
- /
- toc {
display: none; }
- mytoc {
background: none; width: 200px; }
.toc { border: 0px solid; }
- toc ul ul,.toc ul ul {
margin: 0 0 0 1em; }
table.toc { background-color: #f0f4f4; }
- mytoc a,#mytoc a:visited {
font-size: normal; color: #222222; }
- mytoc a:hover {
font-color: #009ee0; /* text-decoration: underline; */ }
- wiki-toc {
width: 200px; margin-top: 6px; float: left; }
- wiki-body {
margin-left: 200px; padding-left: 12px; padding-right: 35px; }
- toc #toctitle,.toc #toctitle,#toc .toctitle,.toc .toctitle {
text-align: left; }
- toc h2,.toc h2 {
font-weight: normal; font-size: 17px; }
/*
- wiki-contents A {
color: #00aeef; }
- wiki-contents A:HOVER {
color: #00aeef; }
- /
- toctitle span {
display: none; }
/*
- wiki-body p,#wiki-body li,#wiki-body dd,div.thumbcaption {
font-size: medium; }
- /
/* required to avoid jumping */
- tocScrolWrapper {
/* left: 450px; */ position: absolute; /* margin-left: 35px; width: 280px; */ }
- tocScrol {
position: absolute; top: 0; /* just used to show how to include the margin in the effect */ /*margin-top: 20px; */ /* border-top: 1px solid purple; */ /*padding-top: 19px;*/ }
- tocScrol.fixed {
position: fixed; top: 0; }
- editPageTxt {
text-align: left; padding-left: 15px; }
- editPageTxt P {
clear: both; }
- toPageTop {
float: left; position: relative; top: 18px; left: 13px; color: #d13f31; } </style>
<script type="text/javascript"> $(document).ready(function() { var parentTable = $("#toc").parent(); $('#mytoc').append($("#toc").first());
$('#mytoc').find("#toc").attr("id", ""); parentTable.closest('table').remove(); });
$(document).ready( function() { var top = $('#tocScrol').offset().top - parseFloat($('#tocScrol').css('marginTop').replace( /auto/, 0)); var nav = $('#tocScrol'); var max = $('#indexing').offset().top - nav.height();
$(window).scroll(function(event) { // what the y position of the scroll is var y = $(this).scrollTop();
if (y > top) { // && signs are html decoded thus this construction if (y >= max) { nav.removeClass('fixed'); nav.css({ position : 'absolute', top : max - top }); } else { nav.addClass('fixed'); nav.removeAttr('style'); } } else { nav.removeClass('fixed'); nav.removeAttr('style'); } }); }); </script> <html> <html>
<a id="toPageTop" href="#">▲</a>
[<a href="http://openwetware.org/index.php?title=Biomod/2013/Aarhus/</html>Achievements And Future Work<html>&action=edit">edit this page</a>]
</html>
Achievements and future work
Origami
Two DNA origamis were designed: a two-layered base plate and a dome that could function as a lid. These were successfully folded and characterized using gel electrophoresis, AFM and TEM. However, connecting the two origamis was not achieved within the time frame of the BIOMOD competition.
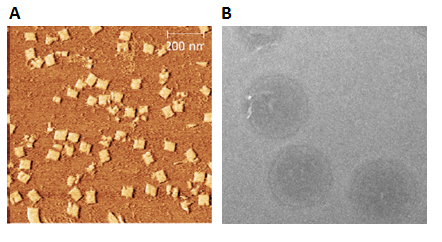
Following the BIOMOD competition more experiments in connecting the plate to the dome will be performed, both using staple strands and the peptide lock. Furthermore, the complete structure with its peptide locks will need testing, to see if it can open in response to cleavage by MMP2, possibly by use of Förster resonance energy transfer (FRET).
Peptide lock
The synthesis of the peptide lock was attempted using a gradual buildup of DNA, but was unsuccesful. Coupling of one DNA strand of the lock to the peptide was achieved using click reactions, but the amide coupling of the amine-modified second DNA strand to the peptide was not achieved. After the BIOMOD competition, amide coupling with standard peptide reagents (HBTU, HOBt, etc.), have been planned to take place in order to obtain the desired molecule. Parallel a design using the succesfull click reaction will be investigated.
Chemical modifications
The photosensitizer was successfully synthesized in five steps from the extracted natural compound and subsequently conjugated to DNA. A cholesterol derivative with an NHS-ester handle was synthetized and subsequently coupled to amine modified DNA strands. Furthermore, to achieve a more direct way of labeling DNA with cholesterols, work will be continued in optimizing the triphosphate synthesis and the following purification. This can be used to synthesize a cholesterol modified nucleoside triphosphate which can be used in a cholesterol labeling-kit with TdT.
System in action
The ability of the photosensitizers to induce apoptosis in cells was investigated and it was shown that photosensitizer-DNA alone could not kill cells, but was functional in complex with a cholesterol-DNA. Future experiments will include expanding the cell analyses to anneal the photosensitizer-DNA conjugate into the origami plate and investigate the ability of this system to induce apoptosis. Cholesterol- and photosensitizer modified DNA strands were successfully incorporated into an origami plate. Cells were treated with the cholesterol-modified plates and visualized using confocal microscopy. As no origamis were visible in these images due to low concentrations of the sample, this experiment will be repeated.
<html></html>
<html> <head> <style>
- indexing {
/* float: left; position: center; */ background-color: #222; border-top: 2px solid #d13f31; color: #006e9c; margin: 0px; padding: 0px 0px 10px 0px; width: 100%; text-align: center; }
.footer-section { padding: 10px; display: table-cell; text-align: left; }
.footer-section-title { font-size: 20px; }
- footer-contents {
color: #006e9c; display: inline-table; }
.footer-section A { color: #006e9c; text-decoration: none; }
.footer-section A:HOVER { color: #00aeef; }
.footer-section ul { list-style-type: square; }
- sitemapTitle {
margin-top: 20px; font-size: 24px; }
- editFooter {
float: right; margin-top: -28px; margin-right: 5px; }
- editFooter A {
color: #006e9c; text-decoration: none; }
.cf:before,.cf:after { content: " "; /* 1 */ display: table; /* 2 */ }
.cf:after { clear: both; }
- bodyContent a[href^="mailto:"], .link-mailto {
background: url() no-repeat scroll right center transparent; padding-right: 0px; color: #006e9c;
}
</style> </head> <body>
SITEMAP | BIOMOD 2013 NANO CREATORS | Aarhus University
Copyright (C) 2013 | BIOMOD Team Nano Creators @ Aarhus University | Programming by: <a href="mailto:pvskaarup@gmail.com?Subject=BIOMOD 2013:">Peter Vium Skaarup</a>.
<img alt="Sigma - Aldrich" src="http://openwetware.org/images/3/39/Sigmaaldrich-logo%28transparant%29.png" width="300px" height="154px"> <img alt="VWR International" src="http://openwetware.org/images/2/28/Vwr_logo.png" width="300px" height="61px"> <img alt="Promega" src="http://openwetware.org/images/7/72/Promega.png" width="175px" height="105px" style="padding-right: 5px; padding-left: 5px;"> <img alt="kem-en-tec" src="http://openwetware.org/images/3/3a/Kementec.png" width="130px" height="129px"> <img alt="Centre For Dna Nanotechnology" src="http://openwetware.org/images/4/4f/CDNA_logo.png" width="420px" height="90px"> <img alt="Dansk Tennis Fond" src="http://openwetware.org/images/9/9a/Dansk_tennis.png" width="250px" height="53px">
</body> </html>