CH391L/S12/Fluorescent Proteins
Green Fluorescent Protein (GFP)
History
GFP was first discovered by Osamu Shimomura in Aequorea jellyfish as a companion protein to the aequorin responsible for the blue glow of the organism.[1][2] Shimomura and his group further characterized and identified the peak luminescence of GFP as similar to that of Aequorea tissue, both of which differed from the peak of the aequorin protein significantly, indicating that GFP altered the color of the aequorin from its natural blue to the green expressed by the organism. They showed that the mechanism for this was transfer of energy from the aequorin to GFP in the presence of a cation[3] The crucial breakthrough came when Douglas Prasher et al cloned the gene and identified its amino acid and DNA sequence.[4] Further characterization showed that expression of the gene led to luminescence in other organism, providing the key inference that all of the information necessary for post-translational synthesis of the chromophore was in the gene itself, and no jellyfish-specific enzymes were needed for production of functional GFP.

Structure and Characterization
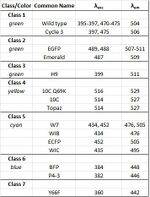
GFP consists of a single β-sheet with alpha helices containing the covalently bonded chromophore 4-(p-hydroxybenzylidene)imidazolidin-5-one (HBI) running through the center. The fluorescence of GFPs is dependent on the key sequence Ser-Tyr-Gly (amino acids 65–67).[5] Wild type GFP has been shown to have 2 excitation peaks at 395-397nm and 470-475nm. The emission spectrum of wild type GFP has a single peak at 504nm.
GFP Derivatives (excluding EGFP and destabilized GFPS)
As of 1998, there were 7 main classes of GFP:
- wild-type mixture of neutral phenol and anionic phenolate (above)
- phenolate anion
- neutral phenol
- phenolate anion with stacked pi electron system
- indole
- imidazole
- phenyl
The excitation and emission wavelength spectra and the color of emission of each of these derivatives are different, as shown in the table. The first 4 classes are polypeptides with a Tyr and position 66, while the final 3 have Trp, His and Phe at that position respectively.[2]
Enhanced GFP (EGFP)
While wild-type GFP revolutionized the observation and monitoring of gene expression, there were several problems associated with it. It's intensity after excitation is low in many cell types, it takes time to fluoresce after protein formation, and it is expressed poorly in multiple mammalian cell types. In order to counter these problems, various types of enhanced GFP's have been created. In one such example, 2 point mutations were made in the GFP chromophore: Ser65 to was changed to Thr and Phe64 was changed to Leu. EGFP was shown to possess many favorable characteristics over not only GFP, but also other forms of expression. Firstly, it requires no cofactors, enzymes, or additional gene products to fluoresce, moving it past non-GFP gene markers immediately. In comparison to GFP, it fluoresces 35 times brighter when excited by blue light, is more soluble, and has more efficient protein folding characteristics than wild-type GFP. In addition, the altered amino acids in EGFP are more prevalent and preferred in eukaryotic cells.[6]
Destabilized GFP
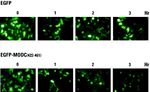
GFP is extremely useful due to its stability; however, this same stability limits the applications of GFP in studies that require rapid turnover of the reporter. In order to compensate for this, various forms of destabilized GFP have been created. Amino acids 422-461 of the degradation domain of mouse ornithine decarboxylace (MODC) were added to the C-terminus of EGFP that degrades in the presence of cycloheximide (CHX). This sequence, known as the PEST sequence, has been linked with protein degradation due to correlation in C-terminal sequences of proteins with shot half-lives. Modification of the PEST sequence added onto EGFP led to various rates of degradation of the protein. The image on the right shows that in the presence of CHX, the modified EGFP (EGFP-MODC) degraded rapidly in comparison to standard EGFP.[5]
Split GFP
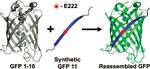
Split GFP's are GFP polypeptides that have been separated and can be re-formed in vivo. Together, the two form the complete GFP protein and fluoresce with an intensity that can be measured. The most commonly used form of split GFP consists of a "superfolder" GFP that is particularly stable when split into GFP 1-10 and GFP 11. GFP 1-10 the first 214 amino acids of GFP, including residues 65-67 that become the chromophore) and a synthetic GFP 11, which completes the β-barrel of the GFP protein. The separate pieces of GFP can be bound to proteins that interact; when interaction occurs, the GFP fragments spontaneously reassemble and fluoresce, allowing for in vivo measurement of protein interaction. Split GFP is particularly helpful because it has increased solubility relative to standard EGFP, and GFP 11 is so small that it can be bound to proteins without affecting their function greatly.[7]
Luciferase
Luciferases are catalysts of enzymatic reactions that cause emission of light in the visible spectrum (bioluminescence).[8]
Bacterial Luciferases
Bacterial luciferases exist mostly in marine Gammaproteobacteria and have various purposes and requirements. One of the cooler functions of bioluminescent bacteria is in symbiotic relationships with higher organisms, such as that of the bacterium in the specialized organism of the angler fish.
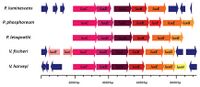

The most well known bacterial systems are the lux systems of the Photobacterium phosphoreum, P. leiognathi, Vibrio harveyi, V. fischeri, and P. luminescencs. All of these systems are basically encoded by the same luxCDABE system, with variations (image on the right). The A and B genes code for the α and β subunits of the heterodimeric protein. The flanking genes code for a fatty acid reductase complex that is required for regeneration of the luciferin fatty aldehyde.
Regulation of Bioluminescence
The LuxI/LuxR system in Gram-negative bacteria is well-understood. luxI encodes a synthase responsible for producing an autoinducing signalling molecule that is expressed constitutively at a low level. Further upstream and in the opposite direction, luxR codes for a transcriptional activator which is also expressed constitutively at a low level. The regulatory region in between the two genes contains the lux box with a binding site for LuxR, which only binds after activation by the molecule produced by luxI. Since these are part of the same operon, the autoinducer triggers its own synthesis, enhancing luminescence in a positive feedback loop. Regulation in the other Vibrio bacterium is far more complex, consisting of various signalling molecules that are autoinducers binding to separate receptors and suppress expression of the lux operon through a variety of steps. AT this point, the regulation of bioluminescence in P. luminscens is not fully understood.
Production of Light
The dimeric enzyme binds directly to a molecule of flavin mononucleide (FMNH2) and catalyzed oxidation of the bound mononucleide and a long-chain aliphatic aldehyde in the presence of molecular oxygen. In the presence of molecular O2, the protein bound to FMNH2 converts it to a peroxyflavin, which reacts with aldehyde to form a new compound, resulting in emission of light at 490nm. Vibrio luciferases have been used to develop a number of reporter vectors, while the P. luminescens lux system is used for imaging of infections and gene expression in mammalian cell lines due to better function at higher temperatures.
Eukaryotic Luciferases
The 2 main eukaryotic systems used in molecular biology are the proteins from the North American firefly P. pyralis and the sea pansy R. reniformis.
P. pyralis
The mechanism by which luminescence is regulated in P. pyralis lanterns is not well understood. The luciferase enzyme is found in peroxisomes of specialized cells called photocytes. The current hypothesis assumes that regulation of oxygen to photocytes is the means of regulation of luminescence. The P. pyralis luc gene encodes a protein which catalyzes the oxidation of D-luciferin to oxyluciferin in 2 steps, a side-effect of which is emission of light of a wavelength 550-570nm. The catalyzed reaction occurs in the presence of molecular oxygen, ATP, and magnesium ions.
R. reniformis
The luciferase present in R. reniformis (Ruc) is triggered by mechanical stimulation, perhaps directly coupled to a neuronal impulse. Stimulus induces uptake of calcium ions into lumisomes in the photocytes, which then triggers a cascade of events that eventually leads to the production of an oxyluciferin compound in the presence of molecular oxygen and the emission of light with a maximum wavelength of 480nm.
Luciferases vs. GFP
Luciferases possess many distinct characteristics that provide distinct advantages over GFP for use as reporter genes in bacterial and eukaryotic environments.
- Shorter half life: the half-life of P. luminescens luciferase is dramatically shorter than that of even destabilized EGFP, and can also be lowered by destabilization of the luciferase protein.
- Efficiency: Luciferase does not need to undergo the same level of posttranslational maturation as GFP and other fluorescent reporters, which takes time and is even slower at 37°C. In addition, while fluorescent reporters can be photo-bleached by excitation, the shorter turnover of bacterial luciferases allows for repetitive measurements at higher temporal resolution.
- Luminescence: Luciferases emit longer wavelengths of light than those of GFP, thus they are higher in energy and can penetrate tissue better - this is especially useful in living animals. Also, unlike fluorescent reporters, luciferase does not have to be excited prior to measurement, eliminating background autofluorescence.
One major disadvantage of luciferase usage is that they can only produce light in the presence of oxygen. However, it has been shown that the gene is expressed in high levels under anaerobic conditions so that if samples are withdrawn, they can be exposed to oxygen and luminescence can be measured.
References
- SHIMOMURA O, JOHNSON FH, and SAIGA Y. Extraction, purification and properties of aequorin, a bioluminescent protein from the luminous hydromedusan, Aequorea. J Cell Comp Physiol. 1962 Jun;59:223-39. DOI:10.1002/jcp.1030590302 |
- Tsien RY. The green fluorescent protein. Annu Rev Biochem. 1998;67:509-44. DOI:10.1146/annurev.biochem.67.1.509 |
- Morise H, Shimomura O, Johnson FH, and Winant J. Intermolecular energy transfer in the bioluminescent system of Aequorea. Biochemistry. 1974 Jun 4;13(12):2656-62. DOI:10.1021/bi00709a028 |
- Prasher DC, Eckenrode VK, Ward WW, Prendergast FG, and Cormier MJ. Primary structure of the Aequorea victoria green-fluorescent protein. Gene. 1992 Feb 15;111(2):229-33. DOI:10.1016/0378-1119(92)90691-h |
- Li X, Zhao X, Fang Y, Jiang X, Duong T, Fan C, Huang CC, and Kain SR. Generation of destabilized green fluorescent protein as a transcription reporter. J Biol Chem. 1998 Dec 25;273(52):34970-5. DOI:10.1074/jbc.273.52.34970 |
- Zhang G, Gurtu V, and Kain SR. An enhanced green fluorescent protein allows sensitive detection of gene transfer in mammalian cells. Biochem Biophys Res Commun. 1996 Oct 23;227(3):707-11. DOI:10.1006/bbrc.1996.1573 |
- Kent KP, Childs W, and Boxer SG. Deconstructing green fluorescent protein. J Am Chem Soc. 2008 Jul 30;130(30):9664-5. DOI:10.1021/ja803782x |
- Waidmann MS, Bleichrodt FS, Laslo T, and Riedel CU. Bacterial luciferase reporters: the Swiss army knife of molecular biology. Bioeng Bugs. 2011 Jan-Feb;2(1):8-16. DOI:10.4161/bbug.2.1.13566 |