Diffusible Signal Oscillator
Last updated January/17/2008
Project status: Active
The Design
This Page will be hevily modified soon I've come with so many modifications of this devise that the information you see now will not be the same tomorrow.
Thanks for the comprehension.
I intended to create an artificial oscillator that would be able to synchronize large population of bacteria.
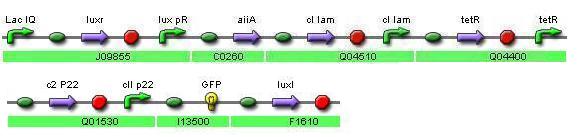
Here I present the second and improved devise that I designed, it still needs refinement, I think I can manage to find a way to assemble it with less biobricks, also I would need to simulate it to adjust the rates for each protein. Still the design have improved since it's predecessor.
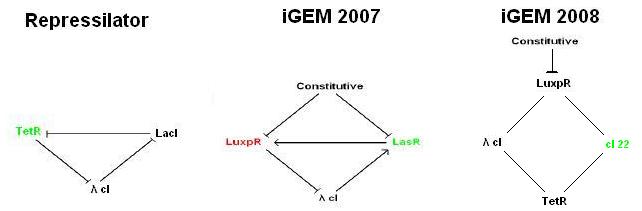
The construction is based upon Ellowitz's Represilator and involves the use of a quorum sensing signal to synchronize the bacteria population.
I suspect that the oscillations can lead to the formation of complex pattens like those formed by drops of water falling on a pond.
In short, I will ask my teammate Luis de Jesús Martínez to model it with the SPIM to see how it works.
Parallel work
- The group of Prakash
A couple weeks before the Jamboree I discovered a very interesting work made at the IAP 2003, a work from the group of Prakash. They basically designed a synchronized oscillator with many similarities with the one I designed. They also remarked a point that I had not noticed "...The half life of HSL is 24 hrs at pH of 7.5 and it is important that a degradation mechanism that is faster than the desired period of oscillations be introduced into the system. Cyclical degration of HSL would generate better synchronisation signals than a constant degradation mechanism."[[1]] They sugested the use of aiiA, an enzime that degrades the lactones, but it seems that the enzyme does not difuse to the medium "The protein has no hydrophobic signal pepetide at the N-terminus and therefore it is believe that it is not secreted. This is supported by the observation that when aiiA is expressed in E.coli DH5alpha or Bacillus 240B1 cells no autoinducer inactivation is detected in the supernatants of these cultures."[[2]] so it seems that lactones will remain there for large amounts of time even with the use of aiiA.
Apparently the work at the IAP 2003 was mainly theoretical but later on, someone actually assembled the constructions that the Pakrash group designed and they are available at the iGEM 2007 kit plates.
BBa_I4204 BBa_I4203 BBa_I4202 BBa_I4201 BBa_I4200
I have analyzed some of them and their design seems incoherent to me... perhaps I just don't understand them well. Anyway, I can't wait to recover those constructions and test them.
- The Group of McGill
I was very surprised to find out that another iGEM team was developing a two phase synchronized oscillator. Like us, they were unable to assemble the whole thing so they only had theoretical work, they say that their construction also produces oscillations... I have my doubts.
They also found the same problem with the degradation of lactones and they also used aiiA.
Further Work
The half life of lactones still bothers me, with aiiA, I could inhibit their effect but I'm afraid that lactones will remain in the medium for a long time. One solution could be the use of peptide signals like those used by Bacillus subtilis, that has been proposed before [[3]] but the system has not been developed and probably it would be to costly for me. The other solution would be to translate the whole construction to an eukaryotic organism and add an excretion signal to aiiA... I have never worked with any eukaryotic organism, perhaps I should start with yeast.
For now I will just develop my skills at the lab, I will need aproval for a procedureto transform the constructions made by Pakrash and see how they work. Also I would like to assemble the Represilator just to practice my assembling skills but for now we still don't have SpeI, so I guess I will have to wait, that bothers me a lot.
I'm afraid that the construction might be to big for a plasmid so that the plasmid would not be able to replicate along with the bacteria. Some days ago I found a biobrick that is 3337bp! I should test it asap but for that I would need approval for a procedure.
I have to study more the Lux promoter... It's function is still alien to me. I fear that aiiA inactivates the lactones before enough cl λ is produced to block the cl promoter... in that case I an put aiiA under the action of cl.
Note: We should build a lux protein generator with R0063. We could score some points by making that part and it would be extremely useful.