Digestion Guide
Which Sites to Digest?
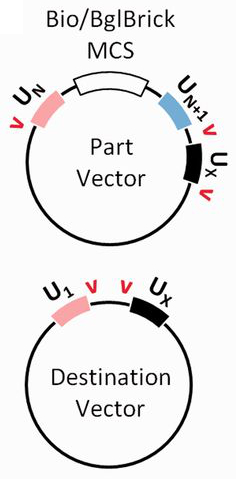
Once DNA sequences of interest are cloned into the part vectors (e.g. by restriction cloning into BioBrick sites in the MCS), UNS-flanked parts can be generated by restriction digestion of sites flanking the UNSes (schematic, right). For all but the most 3' part to be assembled, this involves cutting around the UN and UN+1 sites. The most 3' part should instead be digested around the UN and UX sites. This mixture of parts can then be Gibson assembled with one another and with the U1 and UX sites in each destination vector.
For example, let's say we've cloned genes A, B and C into part vectors U1U2, U2U3 and U3U4. In a three-part assembly, the U1U2 part vector would be cut around the U1 and U2 sites and the U2U3 part vector would be cut around the U2 and U3 sites. The last part, however (U3U4 vector) would be cut around the U3 and UX sites. This would yield the following linear DNAs:
[U1-A-U2] [U2-B-U3] [U3-C-U4-UX]
Digesting the destination vector as shown in the schematic (top right) will give:
[UX-Backbone-U1]
Which can be circularized with the other parts via Gibson assembly.
Because the last (most 3') part in the assembly must be cut with different restriction enzymes than the other parts, it is important to plan the restriction digests carefully to ensure proper assembly. A table of restriction sites that cut at each site on each of the available part and destination vectors is given below. Note that only ONE enzyme must be used per site. E.g. the U1 site of pJT170 can be cut with AflII or SapI; it does NOT require both.

