Donnelly:Research
Research projects
Insecticide resistance in Anopheles gambiae
Our focus is on the primary sub-Saharan African malaria mosquito Anopheles gambiae and metabolic resistance to pyrethroid, DDT and carbamate insecticides.
Association-mapping studies
Study designs
When a disease mutation arises and spreads, marker loci in close proximity will be co-inherited and so will exhibit linkage disequilibrium until broken down by recombination. Population-based association studies provide the potential to detect fine-scaled patterns of linkage disequilibrium (LD), particularly within large outbred populations, thus allowing relatively precise localisation of loci of interest. We are conducting association studies within natural populations (currently Cameroon, Ghana and Uganda) using SNP markers, which are extremely abundant in the Anopheles genome, to evaluate a large panel of insecticide resistance-implicated candidate loci.
SNP discovery1
We sequenced over 300 kilobases across 660 distinct amplicons of the An. gambiae genome using separate pools of M- and S-form An. gambiae as template. We identified a total of 7062 polymorphic features comprising 6995 SNPs and 67 indels, with, on average, a SNP /<50bp. The pooled template sequencing approach proved successful with very low false positive rates and higher false negative rates, but these mostly resulting from low frequency SNPs. SNPs have been submitted to GenBank and will be incorporated into a future version of the Vectorbase Agam annotation.
Screening
SNPs located within and around approximately 260 loci with a known or plausible function in insecticide detoxification(metabolic resistance) or insecticide target-site insensitivity are being screened in a custom-designed Goldengate assay using our Illumina Beadstation 500GX. In addition to candidate SNPs the 1536-SNP assay we are using contains approximately 300 control SNPs to detect and correct for population stratification. In addition to screening of individuals in the Goldengate assay we are also exploring optimum methods for screening pooled templates, which in addition to reducing costs for Illumina consumables, also removes the requirement for whole genome amplification of individual templates2.
Genotype-phenotype association data
The first analyses of SNPs/genes associated with permethrin resistance in Anopheles gambiae populations from Cameroon and Ghana have now been published3 together with a description of the patterns of LD associated with a selective sweep of an insecticide resistance locus 4.
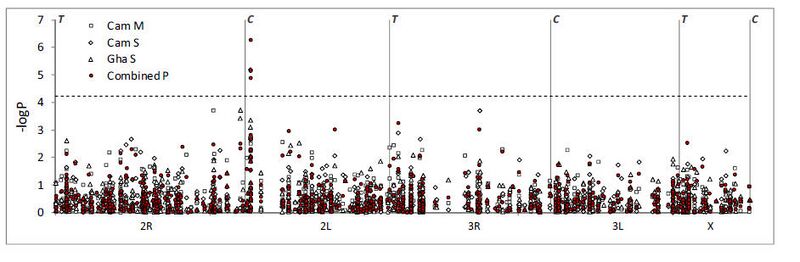
Figure. Association test results for each subsample and for P-values combined across populations (Fisher’s method). The x-axis is a linear physical scale across each chromosome, with centromeres and telomeres denoted by C and T, respectively. The dashed horizontal line shows the P-value level adjusted for multiple testing by Bonferroni correction of the nominal critical α=0.05. Three SNPs, all in the 2L voltage-gated sodium channel, are significantly permethrin resistance phenotypeassociated following strict Bonferroni correction.
Associated papers
1. Wiilding, C.S., Weetman, D., Steen, K., and Donnelly, M.J. (2009) High, clustered, nucleotide diversity in the genome of Anopheles gambiae revealed by SNP discovery through pooled-template sequencing: implications for high-throughput genotyping protocols. BMC Genomics 10, e320
2. Wilding, C.S., Weetman, D., Steen, K., and Donnelly, M.J. (2009) Accurate determination of DNA yield from individual mosquitoes for population genomic applications. Insect Science 16, 361-363
3. Weetman, D., Wilding, C.S., Steen, K., Morgan, J.C., Simard, F. and Donnelly, M.J. (2010) Association mapping of insecticide resistance in wild Anopheles gambiae populations: major variants identified in a low-linkage disequilbrium genome. PLoS One 5, e13140
4. Lynd, A., Weetman, D., Barbosa, S., Yawson, A.E., Mitchell, S., Pinto, J., Hastings, I. and Donnelly, M.J. (2010) Field, genetic and modelling approaches show strong positive selection acting upon an insecticide resistance mutation in Anopheles gambiae s.s. Molecular Biology and Evolution 27, 1117–1125
Gene expression studies
To identify putative genes conferring metabolic resistance we use a microarray-based approach as a screening tool to measure differential expression between susceptible and resistant mosquitoes (Figure 1). Once we have a set of “candidate” genes we screen additional samples using multiplex qPCR. Whilst microarray allows us to obtain a snapshot of the expression profiles of many genes at once the process is time-consuming and expensive. Therefore, we use an alternative method to follow up interesting genes using multiplex PCR allowing us to screen many samples at once though fewer genes (up to 30 per reaction). Once our candidate genes are validated through association mapping and enzyme characterisation (Figure 2) we intend to do further validation by expressing those genes in susceptible mosquitoes to test whether these genes genuinely confer resistance.
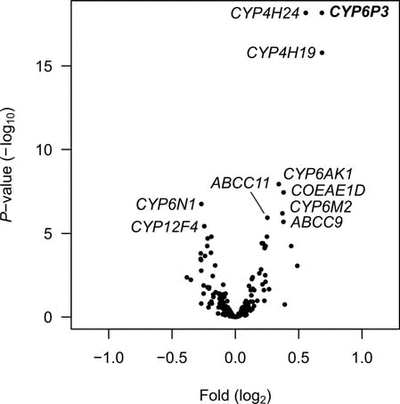
Figure 1. Microarray analysis of loci showing differences in expression levels between permethrin selected and unselected specimens. Each dot represents the mean estimates, P-value vs. fold difference for one unique probe on the microarray. Names are given for the 10 statistically most significant genes.
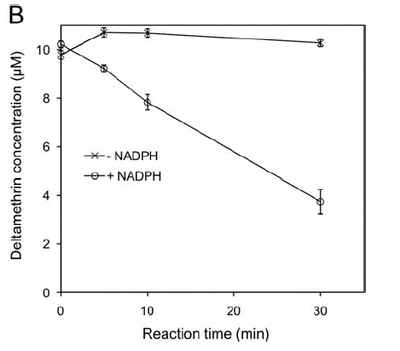
Figure 2. Showing the degradation of the pyrethroid deltamethrin by a recombinant cytochrome P450 Cyp6P3 (see Figure 1).Reactions were performed at 30°C with 10 mM deltamethrin. Concentrations were determined by HPLC peak integration (courtesy Mark Paine).
Associated papers
Müller, P., Warr, E., Stevenson, B.J., Pignatelli, P.M., Morgan, J.C., Steven, A., Yawson, A.E., Mitchell, S.N., Ranson, H., Hemingway, J., Paine, M.J.I., and Donnelly, M.J. (2008) Field-caught permethrin-resistant Anopheles gambiae over-express a cytochrome P450 that metabolizes pyrethroids. PLoS Genetics 4, e1000286
Molecular genetics of tsetse flies
Tsetse flies (genus Glossina) are the major vectors of human and African trypanosomiasis in sub-saharan Africa. Vector control strategies rely on knowledge about the vector. This project focuses on using molecular and morphological approaches to understand how many, what and where are the vector species, especially within the riverine (palpalis group) tsetse flies. This has allowed the design of a simple PCR based species diagnostic for morphologically similar species.
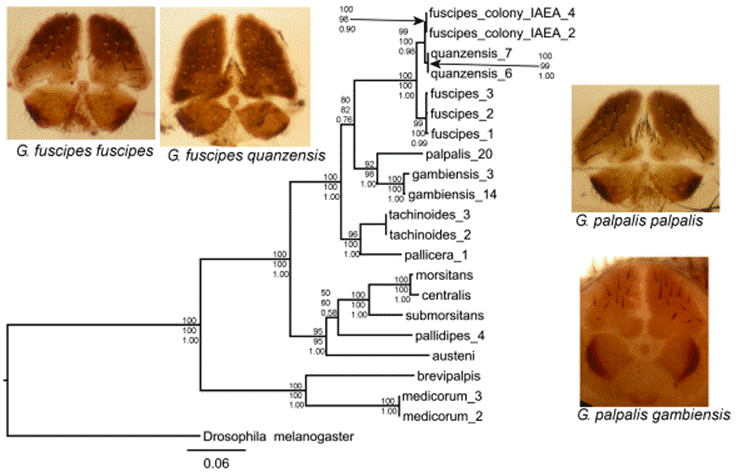
Figure 1. A Glossina phylogeny based on three mitochondrial genes. The photos show the female genital plates of four palpalis group species. The tree shown is the consensus of trees constructed using Maximum Likelihood, distance and Bayesian methods. Bootstrap support for branches for each method is shown at the nodes.
Using population genetics as a tool, we are involved in projects to direct vector control in Senegal, Equatorial Guinea and Guinea. Genetically isolated populations can be targeted for elimination, whereas populations which are not isolated are more suitable for area wide control.
In collaboration with Sophie Ravel (Laboratoire de recherches et de coordination sur les Trypanosomoses, IRD/CIRAD, France) the specific status of three G. fuscipes subspecies (fuscipes, martinii and quanzensis) is being investigated using microsatellites, mitochondrial DNA and the endosymbiont Wigglesworthia glossinidia
Evolution of the epsilon class Glutathione-S-Transferases (GSTs) in Anopheles
GSTs are phase II metabolizing enzymes that detoxify compounds by catalyzing the conjugation of xenobiotics activated in phase I to a water-soluble substrate. The epsilon class is insect specific and has been associated with resistance to chemical insecticides. Using a primer walking approach we have identified and characterized a cluster of epsilon GST genes in species of Anopheles that are from different sub genera and/or inhabit different ecological niches: An. stephensi, An. funestus and An. plumbeus and compared these to their orthologues in An. gambiae. The GST cluster provides an excellent model to investigate gene structure, gene expression and adaptative evolution and the study of its structure and function in Anopheles species is of potential practical interest and may provide insight on the functional diversification of this gene family in insects.
Associated papers
Ayres, C.F.J., Müller, P., Dyer, N.A., Wilding, C.S., Rigden, D.J. and Donnelly, M.J. (2011) Comparative genomics of the anopheline glutathione S-transferase epsilon cluster. PLoS One (in press)