IGEM:Caltech/2007/Project/construct
|
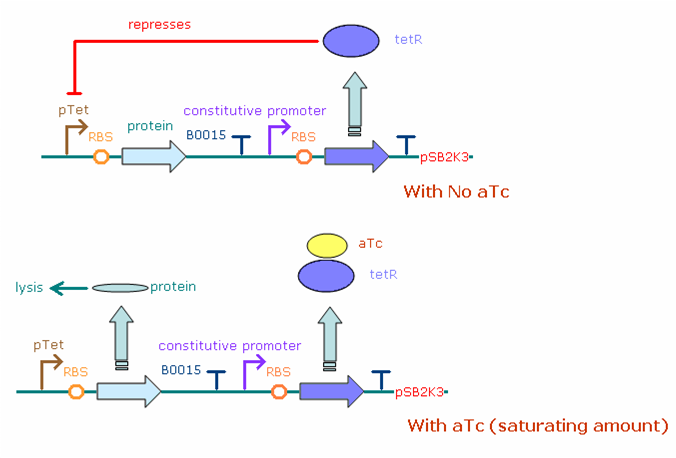
The figure below illustrates the theoretical behavior of the final construct system in E. coli bacteria. The constitutive promoter (both J23100 and J23116) continuously produces transcripts of the tet repressor gene, which when translated turns into tet repressor protein (tetR). In the absence of aTc, the tetR binds to the pTet promoter which drives protein (N/Q/cro/YFP) production, greatly reducing the number of protein transcripts (and thus protein concentration) in the cell. Adding aTc in saturating amounts increases the protein concentration, as aTc binds to tetR, leaving the pTet promoter free to produce more transcripts.
As the cis and trans riboregulators had not yet been designed when the cloning process started, a system with tetR(with parallels to cis repression) and aTc(parallels to trans activation) regulation of protein production was chosen instead, so that verification of lysis could begin on the viral protein constructs. As the recombineered amber phage have not yet been completed, constructs were made in bacterial (D1210) plasmids (pSB2K3). The D1210 strain was chosen for its extremely low-copy number, as was the plasmid (only inducible with IPTG), thus further simulating future infection conditions, where only one phage would infect each cell (each cell hopefully has only one copy of the current plasmid, through the combined choices of D1210 and pSB2K3). |