IGEM:Harvard/2006/Adaptamers/Notebook/2006-7-31
7/31
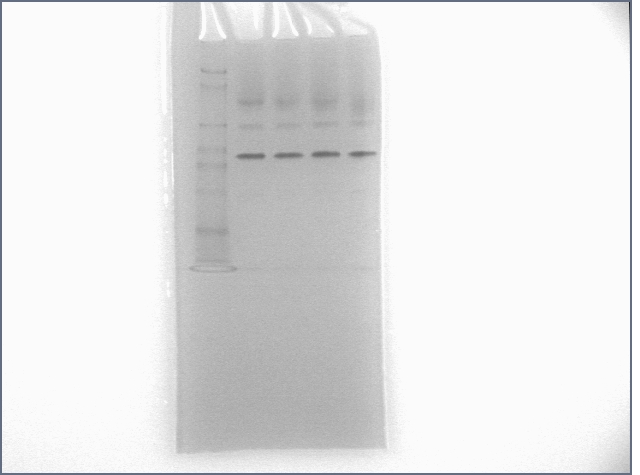
Ran pure thrombin aptamer sequence in a gel along with some friends for one hour. Results below.
Lanes:
1: protein ladder
2: thrombin
3: thrombin + T0
4: thrombin + T5
5: thrombin + T35+S35
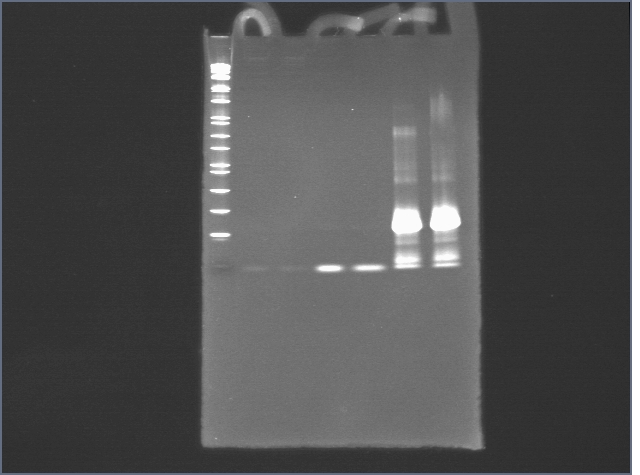
6: DNA ladder
7: T0
8: T0 + thrombin
9: T5
10: T5 + thrombin
11: T35+S35
12: T35+S35 + thrombin
Protein: 20 pmol; DNA: 40 pmol. T35+S35 incubation: 30 minutes before addition of thrombin. 12% Polyacrylamide gel run for 1 hour at 120V.
If you compare lanes 2 with 5 and 11 with 12, you will notice a gel shift. If you were here, you'd notice me doing a little dance. :) :) :)
I should curb my enthusiasm a bit and note that the major thrombin band did not shift. Alain suggested that this might be a reason that papers tend to label the DNA, not the protein.
T35+S35 must be heavy enough to not run too far away while the aptamer region is coming off thrombin, unlike T0 (the pure thrombin aptamer). Running the gel for 1 hour may have made a difference, too. Maybe we'll see an even better shift using T50+S50.