IGEM:Harvard/2006/Cyanobacteria/Notebook/2006-7-7
<html><style type='text/css'> .tabs {
font-size:80%;
font-weight:none;
width: 100%;
color: #FFFFFF;
background:#FFFFFF url("/images/5/54/DarkgreenTab-bg.gif") repeat-x bottom;
}
.tabs li {
background:url("/images/3/36/DarkgeenTab-left.gif") no-repeat left top;
}
.tabs a,.tabs strong {
background:url("/images/d/d3/DarkgreenTab-right.gif") no-repeat right top;
color:#FFFFFF;
padding: 3px 10px 3px 4px;
}
.tabs strong{
color:#CCFF00;
background-image:url("/images/b/b1/DarkgreenTab-right_on.gif");
}
.tabs a:hover{
color:#66FF00;
}
</style></html>
Modeling
More alpha functions to write:
Sinusoidal
What it sounds like
Square pulse
Maybe
Feedback
Feedback is modeled with a couple simple equations: (K_m = Michaelis-Menten)
- Promotion: [math]\displaystyle{ \alpha([P]) = \frac{[P]}{k_m + [P]} }[/math]
- Repression: = [math]\displaystyle{ \alpha([P]) = \frac{k_m}{k_m + [P]} }[/math]
A new hope
The new PCC 7942 culture we ordered finally arrived today!
Growing strains
Liquid culture
Based on the handout which came with the shipment... we created 2 liquid cultures, each in a 500mL earlemeyer flask. One had 100mL of BG11 w/o thiosulfate, and the other has thiosulfate. Each of the samples has about 500uL of liquid culture; the number is so varied due to the pipet used (needed to be absolutely sterile). The handout recommended 2mL... but that would totally drain our stock. Growing at 30C under LL.
Solid culture
Using new yellow cell streaks; streaked two plates, put in incubator.
Original stock
Still has a little left; parafilmed and left at RT at bench. Recommended 26C; 2000-3000lux.
PCR part 2
This PCR is a retry of the PCR done yesterday, except this time we are going to use the liquid live stock as the template instead of the dead cells yesterday, which did not work (why are we not suprised? :D).
The template was formed from ~50uL of stock lysed @95C for 5 min in 100uL of H20. It is labeled "PCC7942" in the freezer.
3 Samples were made: 2 experimental and 1 control. The experimental had:
5uL template 5uL 10x Buffer 1uL dNTP (10mM ea) 1uL Primer (extF) 1uL Primer (extR) 0.5uL HotStarTaq 36.5uL dH20
The run cycle was:
#94C, 15' #94C, 30" #56C, 30" #72C, 1.5' #Cycle to step 2, 7x #94C, 30" #65C, 30" #72C, 1.5' #Cycle to step 6, 30x #72C, 5' #4C hold
In retrospect the 1.5' extension time is not enough... as it is ~3kb the time should be more like 3 minutes.
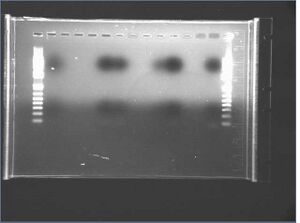
The gel is 1% Agarose, run @120V @ 45m.
Result: =(