IGEM:Harvard/2006/DNA nanostructures/Notebook/2006-7-23
From OpenWetWare
Jump to navigationJump to search
Nanostructure-streptavidin binding experiments
Because purification/precipitation experiments have been unsuccessful, and I don't want to waste any more time, I increased the amount of streptaviding we'd been planning to run by 10x and ran it anyway.Matthewmeisel 19:57, 23 July 2006 (EDT)
- goals:
- run purified DNA nanostructures on PA gel to determine its gel motility
- run scaffold and streptavidin together on PA gel to show that there is no scaffold-streptavidin interaction (we hope!)
- run concentrated, purified DNA nanostructures on a PA gel, with and without streptavidin, with biotin sites facing inwards and outwards, with and without lid latches that are added before streptavidin incubation
- lanes to be run: (total volume of each: 17 μL)
Pre-working stocks
| lane | streptavidin (2 μM) (μL) | p7308 scaffold (44 nM) (μL) |
oligos (100 nM) (μL) | nanostructures (~10 nM in nanostructures, ~90 nM in unused oligos) (μL) |
water (μL) | 1x DNA ladder in 1x TBE/glycerol loading dye (μL) | 10x Tris-gly loading dye (μL) |
| 1 | - | - | - | - | 7 | 10 | - |
| 2 | 2.5 | - | - | - | 12.5 | - | 2 |
| 3 | - | 11.4 | - | - | 3.6 | - | 2 |
| 4 | 2.5 | 11.4 | - | - | 1.1 | - | 2 |
| 5 | - | - | 5.0 (3.2.7.2b) | - | 10 | - | 2 |
| 6 | 2.5 | - | 5.0 (3.2.7.2b) | - | 7.5 | - | 2 |
| 7 | - | - | - | 10.0 4.0.Eb | 5 | - | 2 |
| 8 | 2.5 | - | - | 10.0 4.0.Eb | 2.5 | - | 2 |
| 9 | 2.5 | - | - | 10.0 4.0.Ib | 2.5 | - | 2 |
| 10 | - | - | - | 10.0 4.0.Db | 5 | - | 2 |
| 11 | 2.5 | - | - | 10.0 4.0.Db | 2.5 | - | 2 |
| 12 | 2.5 | - | - | 10.0 4.0.Hb | 2.5 | - | 2 |
- original plan:
- ladder
- streptavidin (2.5 μL 2.0 μM)
- scaffold (11.4 μL 44 nM)
- streptavidin (1.0 μL 0.5 μM) + scaffold (11.4 μL 44 nM)
- biotinylated oligos (5 μL 100 nM)
- biotinylated oligos (5 μL 100 nM) + streptavidin (1.0 μL 0.5 μM)
- nanostructure with outward-facing biotinylated oligos (10 μL 10 nM 4.0.Eb)
- nanostructure with outward-facing biotinylated oligos (10 μL 10 nM 4.0.Eb) + streptavidin (1.0 μL 0.5 μM)
- nanostructure with outward-facing biotinylated oligos (10 μL 10 nM 4.0.Ib) and latch + streptavidin (1.0 μL 0.5 μM)
- nanostructure with inward-facing biotinylated oligos (10 μL 10 nM 4.0.Db)
- nanostructure with inward-facing biotinylated oligos (10 μL 10 nM 4.0.Db) + streptavidin (1.0 μL 0.5 μM)
- nanostructure with inward-facing biotinylated oligos (10 μL 10 nM 4.0.Hb) and latch + streptavidin (1.0 μL 0.5 μM) (latch added before streptavidin incubation)
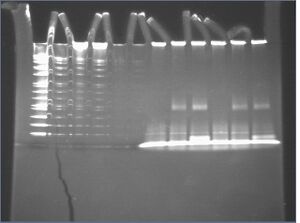

- run on 12% PA gel, 120V at 4[[:Category:{{{1}}}|{{{1}}}]] for 50 min.
- checks at 30 min. and at 50 min. shows that there is no visible dye migration between 30 min. and 50 min.
- stained with 5 μL EtBr in 100 mL water
- assumptions:
- the incubation conditions required for successful streptavidin-biotin are essentially that the two species need to be mixed at room temperature and that the required incubation time is no longer than a few minutes
- results:
- streptavidin-bound DNA nanostructures show no gel shift (lanes 7-12)
- no streptavidin is visible where DNA nanostructures are located at the top of the gel (lanes 8-9, 11-12)
- probably because streptavidin concentration is too small
- streptavidin-bound oligos shift upward (lanes 8-9, 11-12)