IGEM:Imperial/2010/Detection module/Signal Peptide Bearing Protein
The signal peptide bearing protein (SPBP):
In order to set off our signalling cascade in the presence of the cercaria, we have to design a protein that carries our signal peptide out of the cell where the protease has access to it. The protease can then proceed to cleave the peptide (or multiple peptides) off the protein, allowing quorum sensing to take place. One big problem we have to overcome is the cell wall that will obstruct the protease’s access to the SPBP.
Candidate Proteins: CwlB and CwlC
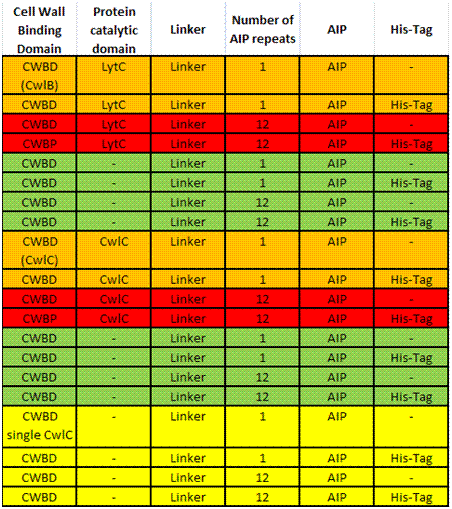
- What are our candidate genes?
The two most promising proteins we have identified are CwlB and CwlC, both of which found naturally on the cell wall of Bacillus subtilis. CwlB, also known as LytC, is a major autolysin. It is involved in cell wall turn-over and more specifically is responsible for break down of peptidoglycans (N-acetylmuramoyl-L-alanine amidase). Cells producing CwlB are uniformly coated with CwlB ater the middle of the exponential phase, which supports the suggestion that CwlB is not associated with cell separation. CwlC is also a hydrolase involved in peptidoglycan break down, however it role is less well understood although it too acts as N-acetylmuramoyl-L-alanine amidase playing an important role in mother-cell lysis during sporulation. Yamamoto et al. Both proteins have in common that they use a cell wall binding domain (CWB) consisting of several repeats to bind non-covalently to the bacterial cell wall using ionic interactions. The CWBs themselves do not show sequence homology and are of very different sizes, but as they function in a similar manner, both can be eluded from the cell wall by solution with a high osmolarity. Mishima et al.
- Why do they should work?
Several papers, especially those by Kobayashi et al., have demonstrated that peptides fused the cell wall binding proteins are expressed correctly and localized on the surface of the cell or alternatively that proteins fused to the CWB of CwlB and CwlC are expressed successfully and localized on the cell wall whilst maintaining the functionality.
Tsuchiya et al. 1999 Kobayashi et al. 2000 Kobayashi et al. 2002 Yamamoto et al. 2003
Additionally, since we do not intend to knock out the original gene from the B. subtilis strain we will not interfere with it normal expression, for example expression at the wrong time, and since we aim at removing the catalytic domain of the protein our fusion protein should not carry out the normal functions, other than cell wall binding, than the original CwlB or CwlC proteins.
- What is the structure of our construct?
First of all we need to add a CWB as this is essential for localization to the cell wall. The is the option to reduce the size of the CWB as it consists of several repeats all but one of which could be removed, however this has not been tested before and might thus not work. Adding the catalytic domain of the protein might help correct localization, but studies have shown that it is not necessary and furthermore activity of the protein at the wrong time might be harmful to the organism. So ideally we want to remove this domain from our construct. Next we add a linker to connect the CWB (or possibly the catalytic domain) to the signal peptide. This linker could either be a simple glycine repeat or, like Yamamoto et al. suggested, this sequence: SRGSRA (for ClwB). In order to get a particularly strong signal we considered using several repeats of the signal peptide (Auto Inducing Peptide: AIP) however we will probably not do so for several reasons: At the site were elastatse cleaves the AIPs could leave four amino acids on each AIP that could interfere with correct binding to its receptor (one lysine at one end and three other amino acids at the other end), with one AIP per protein only a lysine would be added to the end of the AIP which would have a much smaller impact on the function of the AIP. Additionally the receptor requires only 10ng per ml for detection which, as our calculations indicate should reached easily with just one AIP per molecule.
- How do we get the sequences?
We will use PCR to get sequences out of the B. subtilis genome using primers with extensions that will allow us to link the sequences to other DNA elements.
- How do we test for expression?
In order to be able to detect out protein we want to use a His-Tag which will only be added to test constructs but not the final module as it would most likely interfere with the module’s function. The His-Tag would be added to the end of the AIP which probably inhibits correct detection of the AIP by the two component system comD/E. Using a His-Tag would allow us to test for 3 different, very important things: We can lyse some of our bacterial culture and try to isolate our his-tagged protein form the lysate. Detection in this step would demonstrate successful expression of our fusion protein. Then we can place our bacteria in a solution with high osmolarity. As mentioned this eludes the proteins from the cell wall, which should also work for our fusion proteins. Detection of our protein in this step would demonstrate correct expression as well as localization in the cell wall. Last of all we could place the cells in medium containing the elastase or a protease with the same or very similary specificity. If we were able to pull down the his-tagged peptide then it would demonstrate correct expression, localization and that the cleavage site is accessible to the protease and the AIP can be successfully cleaved off.
- Modelling:
How much protein has to be expressed? Will one AIP be enough or should we attempt repeats?
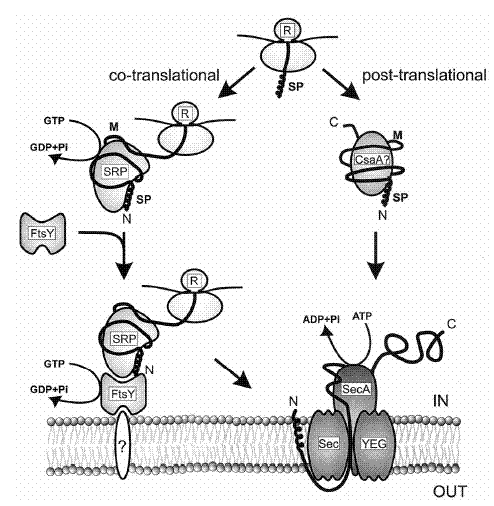
The Secretion System (Sec):
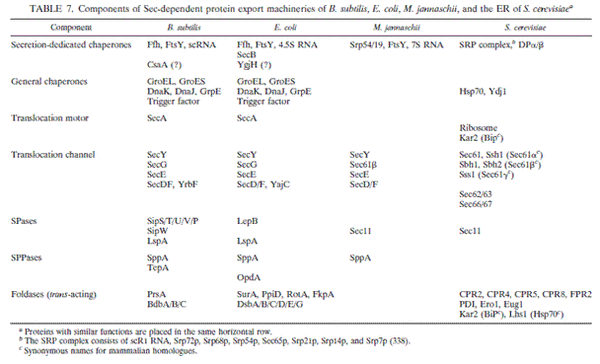
This section, including the diagrams is based on two articles: H. Tjalsma et al. and C. R. Harwood and R. Cranenburgh.The system responsible for secretion of proteins out of the cell seems to be partially conserved across both bacteria and eukaryotes and is called Secretion system or short “Sec”. In B. subtilis the following six components form the sec-dependent secretion machinery:
- Cytosolic Chaperones
The Ffh protein, a GTPase, forms a complex with a small cytosolic RNA forming the Signal Recognition Particle (homologous structures are found in E. coli and S. cerevisiae). It ensures that the protein to be exported remains a secretion competent i.e. linear form by binding to it as it emerges from the ribosome (co-translational export). Unlike in eukaryotes, translation is not arrested until the N-Terminus reaches the membrane but continues as normal. SRP is also thought to be involved in targeting the peptide to the secretion pore and motor. However the protein FtsY plays the most important part in targeting. There is evidence for a second, post-translational, route of protein secretion via CsaA which targets preproteins to SecA. CsaA shows affinity to preproteins and exhibits some chaperone-like activity.
- Translocation motor
In B. subtilis the gene which was originally called div and was identified to be crucial for cell division and sporulation, turned out to be a homologue of E. coli’s SecA protein. SecA is an ATPase responsible for translocation of the preproteins through the translocation channel out of the cell.
- Translocation channel
A heterotrimeric complex of the membrane proteins SecY, SecE, and SecG forms the main core of the translocation channel. SecE and signal peptides bind to the same or overlapping regions in SecY and SecE probably functions as a surrogate signal peptide when the SecY channel is in its closed form in the absence of a translocating protein. When a signal peptide is present, it displaces SecE and allows opening of the channel. SecG is not strictly required for preprotein translocation and cell viability but it is necessary for efficient translocation, possibly by facilitating the movement of preproteins through the translocation channel in concert with the insertion and deinsertion cycles of SecA. Deficiency caused secretion defects that resulted in cold-sensitive growth in B. subtilis as well as E. coli.
Unlike other organisms B. subtilis has a natural fusion protein of two components called SecDF. This protein as well is not essential for cell viability but it is thought the protein regulates SecA activity and might also be involved in translocase assembly and clearing of signal peptides from the channel.
- SPases:
- SPPases:
- Folding factors (on trans site):
Work on the last three section of the secretion system is still in progress.
Transmembrane protein:
Initially we considered a fusion of some transmembrane protein with our signal peptide sequence. There are several problems to consider: The N-terminus cannot be modified as it specifies export via the Sec system, so only the C-Terminus would be available for modification. This implies that a protein would have to be chosen that has an exposed C-Terminus from which the proteases could remove the signal peptide easily. Additionally the cell wall problem would not be solved by this.
Flagellin:
This protein is the most abundant subunit of flagella and makes up most of the structure. The advantages this protein would have to offer is that it is abundantly synthesised and transported is through the cell wall where it would be accessible to the cercarias’ proteases. However structural studies have shown that both C and N-Temini of flagellin are inside the flagellar structure and would thus not be accessible. I believe that modifying the structure of flagellin so that one of the termini is exposed is not feasible and would probably not work as the structure of flagellin is highly conserved across bacterial species, suggesting that deviation from the natural structure is not functional.
Sortase:
Sortases are proteins used by Gram positive bacteria to attach specific proteins to their cell wall. B. subtilis has two of these – YhcS and YwpE – but they are not well characterized. S. aureus however has one – SrtA – which has been tested and is well understood. Sortases recognize a sequence on the C-Terminus of a protein and attach it covalently to the cell wall. About 20 proteins are attached to the cell wall in this way naturally by S. aureus, for example the IG-binding protein A, but other proteins have successfully been attached to the cell wall by creating a signal peptide protein fusion. The sortases themselves are membrane bound via their N-Termini which suggests to me that they will not add the proteins on the outside of the cell wall, however the function of some of these proteins suggests otherwise. Unfortunately I have not been able to determine the localization of the proteins yet. SrtA has a well characterized specificity for LPXTG motifs which it recognizes with its catalytic TLXTC domain near the C-Terminus. Alternatively to SrtA of S. aureus, close relatives to B. subtilis, such as B. anthacis also use sortases of known specificity.
I think this approach is the most promising as it seems best suited to overcome the cell wall problem and also because we might be able to target a synthetic protein (that only contains the secretion sequence on its N-Terminus and the sorting signal at its C-Terminus with many signal peptide-protease cleavage site repeats in the middle) to the cell wall.