IGEM:Melbourne/2008/BioClock/General Modeling Objectives
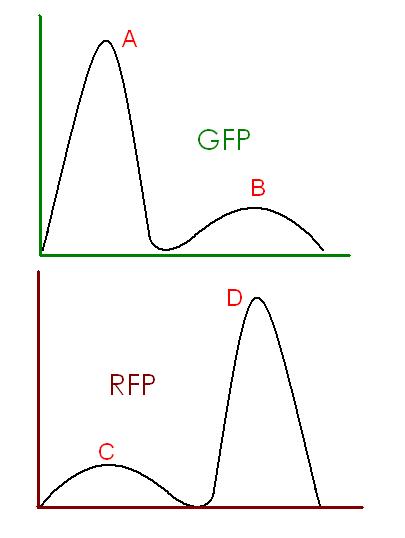
Below is an idealized image of what is outputted from David's version of Stephen's model for which we will try to determine sensitivities.
In order to test for which parameters (kinetic rates/concentrations) are most sensitive/optimal, we will use the following formulas.
[math]\displaystyle{ K = (A-B)/A }[/math]
[math]\displaystyle{ L = (D-C)/D }[/math]
[math]\displaystyle{ W = X/Y }[/math]
Where
[math]\displaystyle{ X = A/C }[/math]
[math]\displaystyle{ Y = D/B }[/math]
Such that values for K, L and W close to one are desirable, meaning that (for K and L) there is a large discrepancy between the supposed off position and on position for the Fluorescent Protein, while W is a measure of the consistency in production of the different proteins. A value of W close to one implies that the visible signal will be distinct for each pulse (i.e, little or no overriding signal from the other protein produced). Similar concentrations of fluorescent proteins are needed for a desirable W value.
If we wish to take into account the different orders of magnitude that fluorescent proteins fluoresce, merely a constant needs to be added:
[math]\displaystyle{ X = (A/C)*(h/j) }[/math]
[math]\displaystyle{ Y = (D/B)*(j/h) }[/math]
where h and j are the constants representing the amount each protein fluoresces under equimolar concentrations.